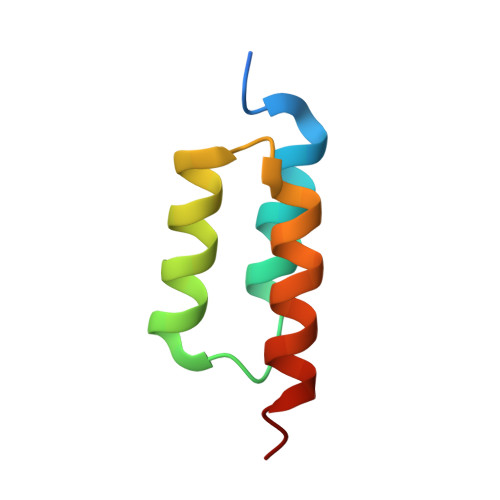
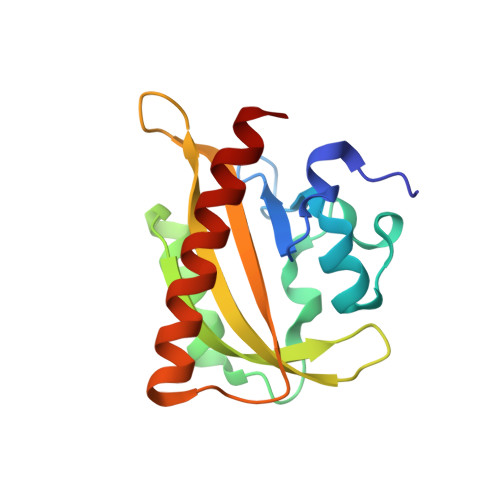
LOVTRAP: an optogenetic system for photoinduced protein dissociation.
Wang, H., Vilela, M., Winkler, A., Tarnawski, M., Schlichting, I., Yumerefendi, H., Kuhlman, B., Liu, R., Danuser, G., Hahn, K.M.(2016) Nat Methods 13: 755-758
- PubMed: 27427858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nmeth.3926
- Primary Citation Related Structures:
5DJT, 5DJU, 5EFW - PubMed Abstract:
LOVTRAP is an optogenetic approach for reversible light-induced protein dissociation using protein A fragments that bind to the LOV domain only in the dark, with tunable kinetics and a >150-fold change in the dissociation constant (Kd). By reversibly sequestering proteins at mitochondria, we precisely modulated the proteins' access to the cell edge, demonstrating a naturally occurring 3-mHz cell-edge oscillation driven by interactions of Vav2, Rac1, and PI3K proteins.
- Department of Pharmacology, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA.
Organizational Affiliation: