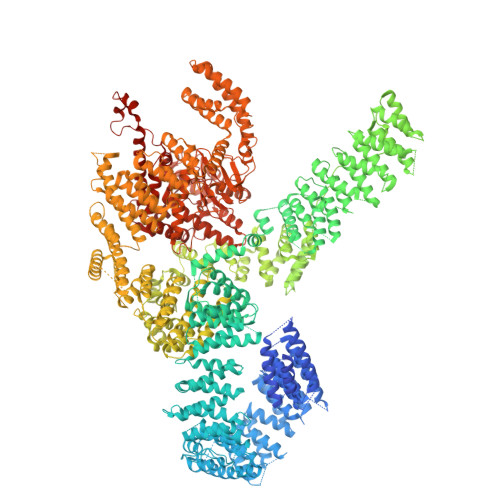
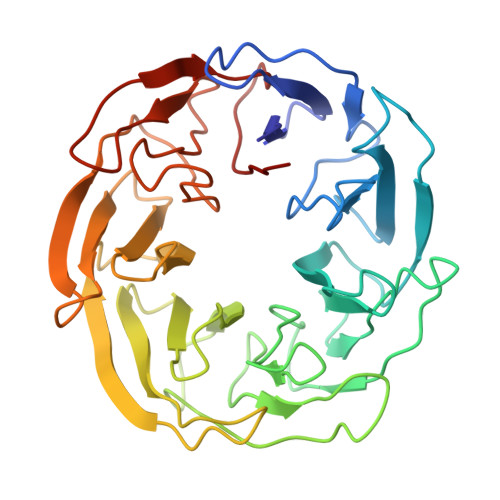
Tor Forms a Dimer Through an N-Terminal Helical Solenoid with a Complex Topology
Baretic, D., Berndt, A., Ohashi, Y., Johnson, C.M., Williams, R.L.(2016) Nat Commun 7: 11016
- PubMed: 27072897 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11016
- Primary Citation Related Structures:
5FVM - PubMed Abstract:
The target of rapamycin (Tor) is a Ser/Thr protein kinase that regulates a range of anabolic and catabolic processes. Tor is present in two complexes, TORC1 and TORC2, in which the Tor-Lst8 heterodimer forms a common sub-complex. We have determined the cryo-electron microscopy (EM) structure of Tor bound to Lst8. Two Tor-Lst8 heterodimers assemble further into a dyad-symmetry dimer mediated by Tor-Tor interactions. The first 1,300 residues of Tor form a HEAT repeat-containing α-solenoid with four distinct segments: a highly curved 800-residue N-terminal 'spiral', followed by a 400-residue low-curvature 'bridge' and an extended 'railing' running along the bridge leading to the 'cap' that links to FAT region. This complex topology was verified by domain insertions and offers a new interpretation of the mTORC1 structure. The spiral of one TOR interacts with the bridge of another, which together form a joint platform for the Regulatory Associated Protein of TOR (RAPTOR) regulatory subunit.
- MRC Laboratory of Molecular Biology, Cambridge CB2 0QH, UK.
Organizational Affiliation: