Structure of the human cohesin regulator Pds5 in complex with Wapl motif
Ouyang, Z., Tomchick, D.R., Yu, H.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sister chromatid cohesion protein PDS5 homolog B | 1,111 | Homo sapiens | Mutation(s): 1 Gene Names: PDS5B, APRIN, AS3, KIAA0979 | 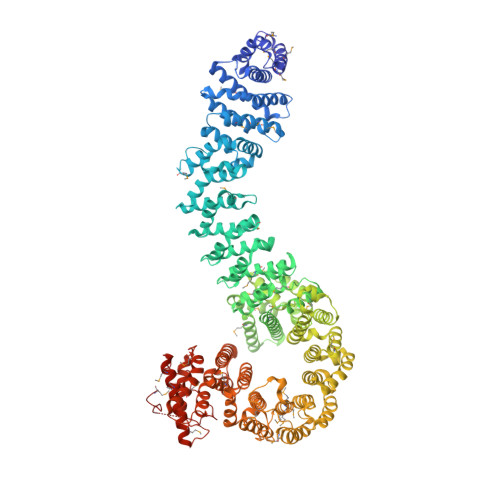 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9NTI5 GTEx: ENSG00000083642 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9NTI5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Wings apart-like protein homolog | C [auth E] | 33 | Homo sapiens | Mutation(s): 0 | 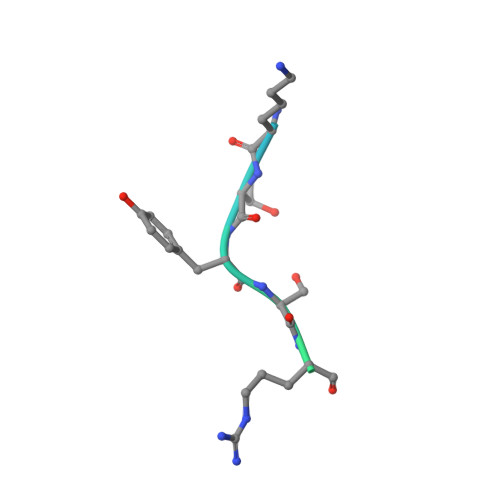 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q7Z5K2 GTEx: ENSG00000062650 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7Z5K2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| IHP Download:Ideal Coordinates CCD File | D [auth A], E [auth B] | INOSITOL HEXAKISPHOSPHATE C6 H18 O24 P6 IMQLKJBTEOYOSI-GPIVLXJGSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 120.76 | α = 90 |
| b = 162.368 | β = 90 |
| c = 173.061 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| PHASER | phasing |