Discovery of (1S,2R,3S,4S,5R,6R)-2-Amino-3-[(3,4-difluorophenyl)sulfanylmethyl]-4-hydroxy-bicyclo[3.1.0]hexane-2,6-dicarboxylic Acid Hydrochloride (LY3020371HCl): A Potent, Metabotropic Glutamate 2/3 Receptor Antagonist with Antidepressant-Like Activity.
Chappell, M.D., Li, R., Smith, S.C., Dressman, B.A., Tromiczak, E.G., Tripp, A.E., Blanco, M.J., Vetman, T., Quimby, S.J., Matt, J., Britton, T.C., Fivush, A.M., Schkeryantz, J.M., Mayhugh, D., Erickson, J.A., Bures, M.G., Jaramillo, C., Carpintero, M., Diego, J.E., Barberis, M., Garcia-Cerrada, S., Soriano, J.F., Antonysamy, S., Atwell, S., MacEwan, I., Condon, B., Sougias, C., Wang, J., Zhang, A., Conners, K., Groshong, C., Wasserman, S.R., Koss, J.W., Witkin, J.M., Li, X., Overshiner, C., Wafford, K.A., Seidel, W., Wang, X.S., Heinz, B.A., Swanson, S., Catlow, J.T., Bedwell, D.W., Monn, J.A., Mitch, C.H., Ornstein, P.L.(2016) J Med Chem 59: 10974-10993
- PubMed: 28002967 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01119
- Primary Citation Related Structures:
5KZN, 5KZQ - PubMed Abstract:
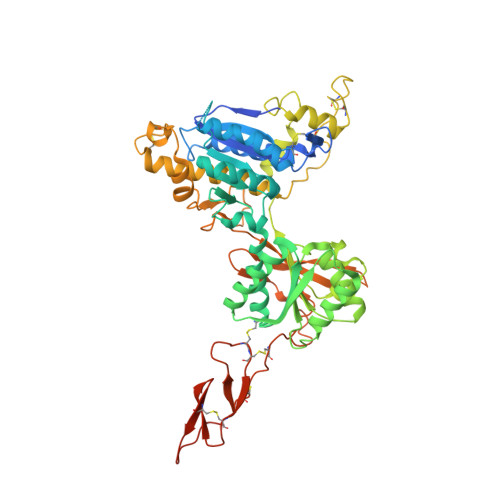
As part of our ongoing efforts to identify novel ligands for the metabotropic glutamate 2 and 3 (mGlu 2/3 ) receptors, we have incorporated substitution at the C3 and C4 positions of the (1S,2R,5R,6R)-2-amino-bicyclo[3.1.0]hexane-2,6-dicarboxylic acid scaffold to generate mGlu 2/3 antagonists. Exploration of this structure-activity relationship (SAR) led to the identification of (1S,2R,3S,4S,5R,6R)-2-amino-3-[(3,4-difluorophenyl)sulfanylmethyl]-4-hydroxy-bicyclo[3.1.0]hexane-2,6-dicarboxylic acid hydrochloride (LY3020371·HCl, 19f), a potent, selective, and maximally efficacious mGlu 2/3 antagonist. Further characterization of compound 19f binding to the human metabotropic 2 glutamate (hmGlu 2 ) site was established by cocrystallization of this molecule with the amino terminal domain (ATD) of the hmGlu 2 receptor protein. The resulting cocrystal structure revealed the specific ligand-protein interactions, which likely explain the high affinity of 19f for this site and support its functional mGlu 2 antagonist pharmacology. Further characterization of 19f in vivo demonstrated an antidepressant-like signature in the mouse forced-swim test (mFST) assay when brain levels of this compound exceeded the cellular mGlu 2 IC 50 value.
- Discovery Chemistry Synthesis Group, Centro de Investigación Lilly S.A. Avda. de la Industria , 30 Alcobendas, Madrid 28108, Spain.
Organizational Affiliation: