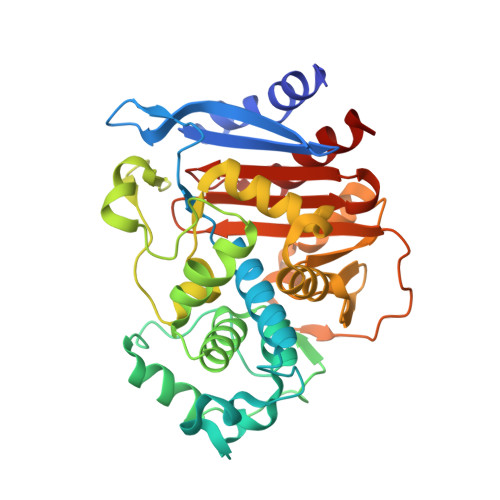
Inhibition of Acinetobacter-Derived Cephalosporinase: Exploring the Carboxylate Recognition Site Using Novel beta-Lactamase Inhibitors.
Caselli, E., Romagnoli, C., Powers, R.A., Taracila, M.A., Bouza, A.A., Swanson, H.C., Smolen, K.A., Fini, F., Wallar, B.J., Bonomo, R.A., Prati, F.(2018) ACS Infect Dis 4: 337-348
- PubMed: 29144725
- DOI: https://doi.org/10.1021/acsinfecdis.7b00153
- Primary Citation of Related Structures:
5W12, 5W13, 5W14 - PubMed Abstract:
Boronic acids are attracting a lot of attention as β-lactamase inhibitors, and in particular, compound S02030 ( K i = 44 nM) proved to be a good lead compound against ADC-7 ( Acinetobacter-derived cephalosporinase), one of the most significant resistance determinants in A. baumannii. The atomic structure of the ADC-7/S02030 complex highlighted the importance of critical structural determinants for recognition of the boronic acids. Herein, to elucidate the role in recognition of the R2-carboxylate, which mimics the C 3 /C 4 found in β-lactams, we designed, synthesized, and characterized six derivatives of S02030 (3a). Out of the six compounds, the best inhibitors proved to be those with an explicit negative charge (compounds 3a-c, 3h, and 3j, K i = 44-115 nM), which is in contrast to the derivatives where the negative charge is omitted, such as the amide derivative 3d ( K i = 224 nM) and the hydroxyamide derivative 3e ( K i = 155 nM). To develop a structural characterization of inhibitor binding in the active site, the X-ray crystal structures of ADC-7 in a complex with compounds 3c, SM23, and EC04 were determined. All three compounds share the same structural features as in S02030 but only differ in the carboxy-R2 side chain, thereby providing the opportunity of exploring the distinct binding mode of the negatively charged R2 side chain. This cephalosporinase demonstrates a high degree of versatility in recognition, employing different residues to directly interact with the carboxylate, thus suggesting the existence of a "carboxylate binding region" rather than a binding site in ADC enzymes. Furthermore, this class of compounds was tested against resistant clinical strains of A. baumannii and are effective at inhibiting bacterial growth in conjunction with a β-lactam antibiotic.
Organizational Affiliation:
Department of Life Sciences , University of Modena and Reggio Emilia , Via Campi 101 , 41125 , Modena , Italy.