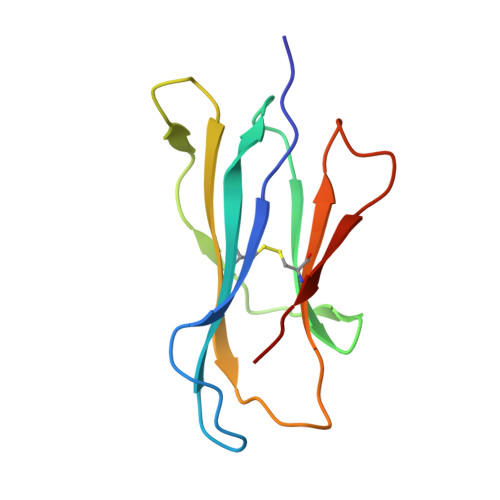
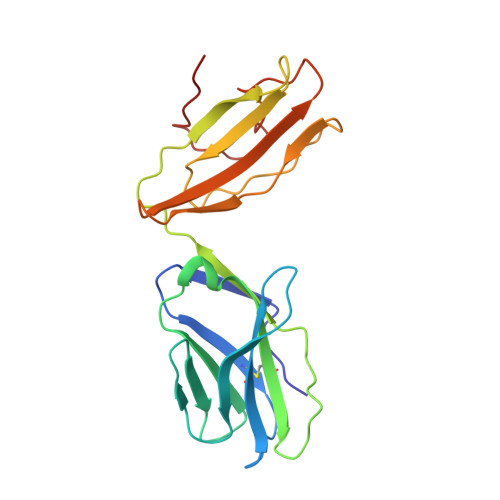
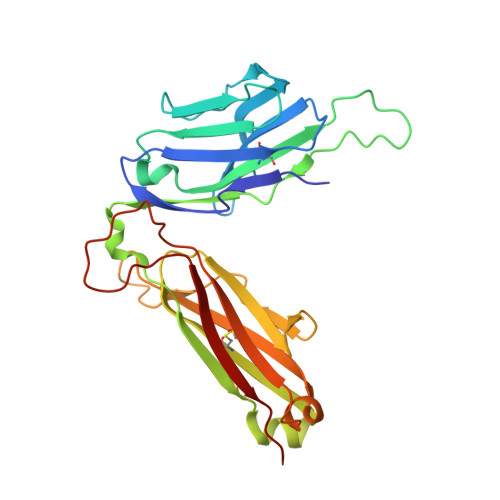
A molecular basis of human T cell receptor autoreactivity toward self-phospholipids.
Shahine, A., Van Rhijn, I., Cheng, T.Y., Iwany, S., Gras, S., Moody, D.B., Rossjohn, J.(2017) Sci Immunol 2
- PubMed: 29054999
- DOI: https://doi.org/10.1126/sciimmunol.aao1384
- Primary Citation of Related Structures:
5WJO, 5WKE, 5WKG, 5WKI, 5WL1 - PubMed Abstract:
Human T cell autoreactivity toward lipid antigens presented by CD1 proteins can manifest in numerous diseases, including psoriasis, contact hypersensitivities, and allergies. However, the molecular mechanisms for regulating T cell autoreactivity toward lipid antigens remain unclear. We determined the basis for T cell receptor (TCR) autoreactivity toward CD1b bound to self-phospholipids. The spectrum of self-antigens captured by CD1b skews toward abundant membrane phospholipids such as phosphatidylcholine and phosphatidylethanolamine. However, TCRs can specifically recognize rare phospholipids, including phosphatidylglycerol (PG). The structure of an autoreactive TCR bound to CD1b-PG shows that discrimination occurs through a marked induced fit movement of PG so that its polar head group fits snugly into the cationic cup of the TCR. Conversely, TCR binding toward ubiquitous self-phospholipids was sterically or electrostatically repelled. Accordingly, we describe a mechanism of TCR autoreactivity toward rare phospholipids and avoidance of autoreactivity to the most abundant self-phospholipids.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: