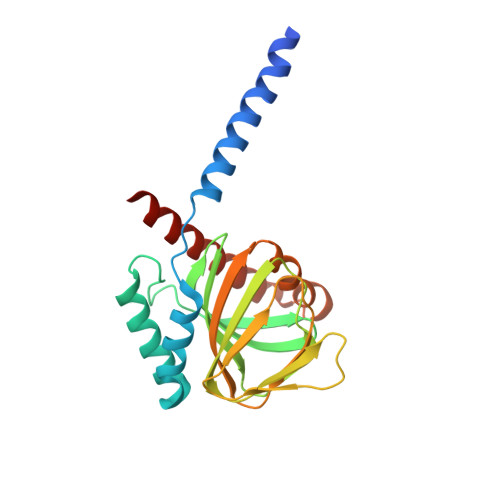
Structural basis for sigma1receptor ligand recognition.
Schmidt, H.R., Betz, R.M., Dror, R.O., Kruse, A.C.(2018) Nat Struct Mol Biol 25: 981-987
- PubMed: 30291362 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-018-0137-2
- Primary Citation Related Structures:
6DJZ, 6DK0, 6DK1 - PubMed Abstract:
The σ 1 receptor is a poorly understood membrane protein expressed throughout the human body. Ligands targeting the σ 1 receptor are in clinical trials for treatment of Alzheimer's disease, ischemic stroke, and neuropathic pain. However, relatively little is known regarding the σ 1 receptor's molecular function. Here, we present crystal structures of human σ 1 receptor bound to the antagonists haloperidol and NE-100, and the agonist (+)-pentazocine, at crystallographic resolutions of 3.1 Å, 2.9 Å, and 3.1 Å, respectively. These structures reveal a unique binding pose for the agonist. The structures and accompanying molecular dynamics (MD) simulations identify agonist-induced structural rearrangements in the receptor. Additionally, we show that ligand binding to σ 1 is a multistep process that is rate limited by receptor conformational change. We used MD simulations to reconstruct a ligand binding pathway involving two major conformational changes. These data provide a framework for understanding the molecular basis for σ 1 agonism.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA, USA.
Organizational Affiliation: