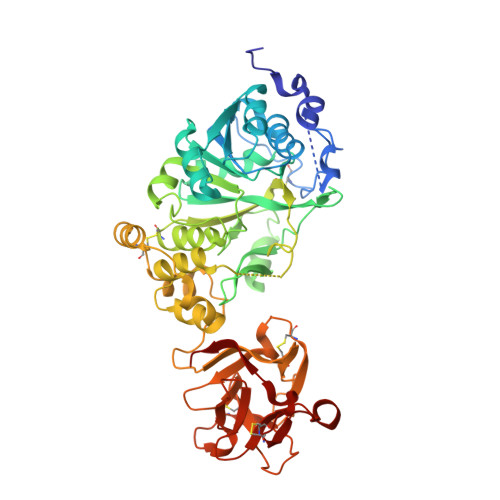
Structural Analysis of a GalNAc-T2 Mutant Reveals an Induced-Fit Catalytic Mechanism for GalNAc-Ts.
de Las Rivas, M., Coelho, H., Diniz, A., Lira-Navarrete, E., Companon, I., Jimenez-Barbero, J., Schjoldager, K.T., Bennett, E.P., Vakhrushev, S.Y., Clausen, H., Corzana, F., Marcelo, F., Hurtado-Guerrero, R.(2018) Chemistry 24: 8382-8392
- PubMed: 29601100
- DOI: https://doi.org/10.1002/chem.201800701
- Primary Citation of Related Structures:
6EGS - PubMed Abstract:
The family of polypeptide N-acetylgalactosamine (GalNAc) transferases (GalNAc-Ts) orchestrates the initiating step of mucin-type protein O-glycosylation by transfer of GalNAc moieties to serine and threonine residues in proteins. Deficiencies and dysregulation of GalNAc-T isoenzymes are related to different diseases. Recently, it has been demonstrated that an inactive GalNAc-T2 mutant (F104S), which is not located at the active site, induces low levels of high-density lipoprotein cholesterol (HDL-C) in humans. Herein, the molecular basis for F104S mutant inactivation has been deciphered. Saturation transfer difference NMR spectroscopy experiments demonstrate that the mutation induces loss of binding to peptide substrates. Analysis of the crystal structure of the F104S mutant bound to UDP-GalNAc (UDP=uridine diphosphate), combined with molecular dynamics (MD) simulations, has revealed that the flexible loop is disordered and displays larger conformational changes in the mutant enzyme than that in the wild-type (WT) enzyme. 19 F NMR spectroscopy experiments reveal that the WT enzyme only reaches the active state in the presence of UDP-GalNAc, which provides compelling evidence that GalNAc-T2 adopts a UDP-GalNAc-dependent induced-fit mechanism. The F104S mutation precludes the enzyme from achieving the active conformation and concomitantly binding peptide substrates. This study provides new insights into the catalytic mechanism of the large family of GalNAc-Ts and how these enzymes orchestrate protein O-glycosylation.
Organizational Affiliation:
Instituto de Biocomputación y Fisica de Sistemas Complejos (BIFI), BIFI-IQFR (CSIC) Joint Unit, Universidad de Zaragoza, 50018, Zaragoza, Spain.