3,5,7-Substituted Pyrazolo[4,3- d]pyrimidine Inhibitors of Cyclin-Dependent Kinases and Their Evaluation in Lymphoma Models.
Jorda, R., Havlicek, L., Sturc, A., Tuskova, D., Daumova, L., Alam, M., Skerlova, J., Nekardova, M., Perina, M., Pospisil, T., Siroka, J., Urbanek, L., Pachl, P., Rezacova, P., Strnad, M., Klener, P., Krystof, V.(2019) J Med Chem 62: 4606-4623
- PubMed: 30943029 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00189
- Primary Citation Related Structures:
6GVA - PubMed Abstract:
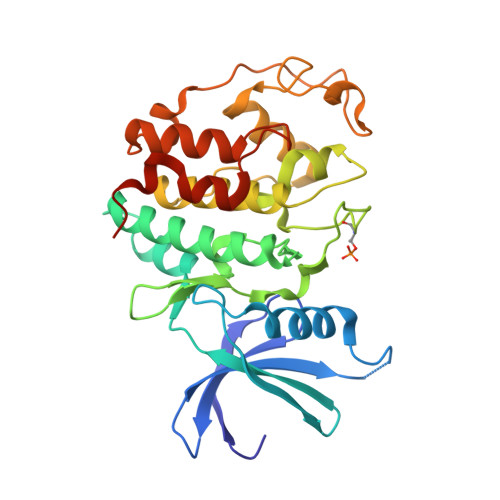
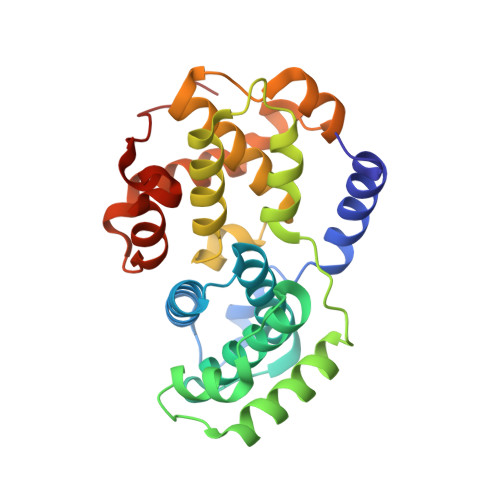
Cyclin-dependent kinases are therapeutic targets frequently deregulated in various cancers. By convenient alkylation of the 5-sulfanyl group, we synthesized 3-isopropyl-7-[4-(2-pyridyl)benzyl]amino-1(2) H-pyrazolo[4,3- d]pyrimidines with various substitutions at position 5 with potent antiproliferative activity in non-Hodgkin lymphoma cell lines. The most potent derivative 4.35 also displayed activities across more than 60 cancer cell lines. The kinase profiling confirmed high selectivity of 4.35 toward cyclin-dependent kinases (CDKs) 2, 5, and 9, and the cocrystal with CDK2/cyclin A2 revealed its binding in the active site. Cultured lymphoma cell lines treated with 4.35 showed dephosphorylation of CDK substrates, cleavage of PARP-1, downregulation of XIAP and MCL-1, and activation of caspases, which collectively confirmed ongoing apoptosis. Moreover, 4.35 demonstrated significant activity in various cell line xenograft and patient-derived xenograft mouse models in vivo both as a monotherapy and as a combination therapy with the BCL2-targeting venetoclax. These findings support further studies of combinatorial treatment based on CDK inhibitors.
- Laboratory of Growth Regulators , Palacký University and Institute of Experimental Botany, The Czech Academy of Sciences , Šlechtitelů 27 , 783 71 Olomouc , Czech Republic.
Organizational Affiliation: