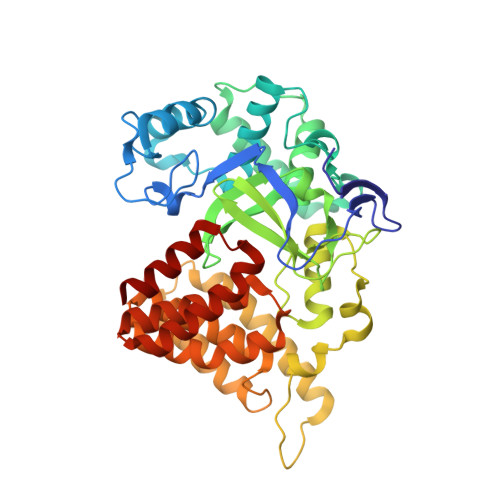
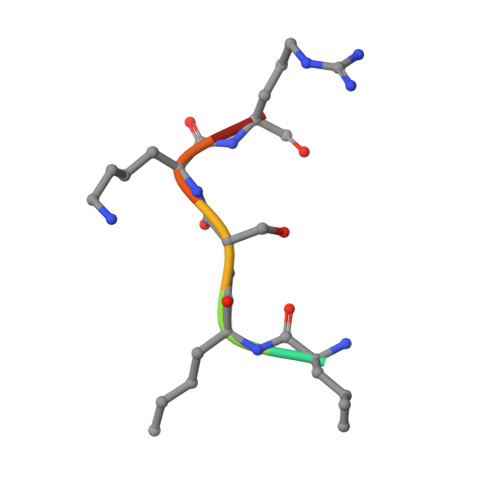
A functional proteomics platform to reveal the sequence determinants of lysine methyltransferase substrate selectivity.
Cornett, E.M., Dickson, B.M., Krajewski, K., Spellmon, N., Umstead, A., Vaughan, R.M., Shaw, K.M., Versluis, P.P., Cowles, M.W., Brunzelle, J., Yang, Z., Vega, I.E., Sun, Z.W., Rothbart, S.B.(2018) Sci Adv 4: eaav2623-eaav2623
- PubMed: 30498785 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aav2623
- Primary Citation Related Structures:
6MON - PubMed Abstract:
Lysine methylation is a key regulator of histone protein function. Beyond histones, few connections have been made to the enzymes responsible for the deposition of these posttranslational modifications. Here, we debut a high-throughput functional proteomics platform that maps the sequence determinants of lysine methyltransferase (KMT) substrate selectivity without a priori knowledge of a substrate or target proteome. We demonstrate the predictive power of this approach for identifying KMT substrates, generating scaffolds for inhibitor design, and predicting the impact of missense mutations on lysine methylation signaling. By comparing KMT selectivity profiles to available lysine methylome datasets, we reveal a disconnect between preferred KMT substrates and the ability to detect these motifs using standard mass spectrometry pipelines. Collectively, our studies validate the use of this platform for guiding the study of lysine methylation signaling and suggest that substantial gaps exist in proteome-wide curation of lysine methylomes.
- Center for Epigenetics, Van Andel Research Institute, Grand Rapids, MI 49503, USA.
Organizational Affiliation: