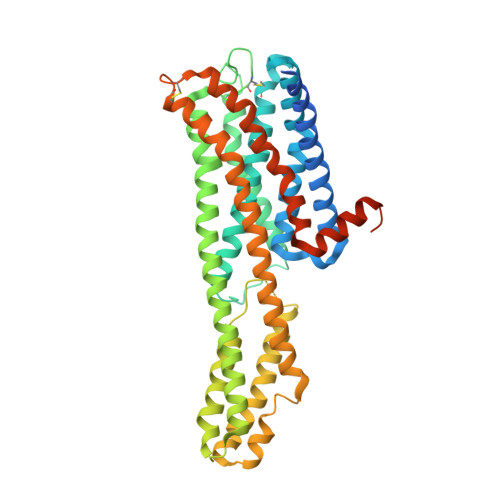
Structure of a Hallucinogen-Activated Gq-Coupled 5-HT 2A Serotonin Receptor
Kim, K.L., Che, T., Panova, O., DiBerto, J.F., Lyu, J., Krumm, B.E., Wacker, D., Robertson, M.J., Seven, A.B., Nichols, D.E., Shoichet, B.K., Skiniotis, G., Roth, B.L.(2020) Cell 182: 1574-1588