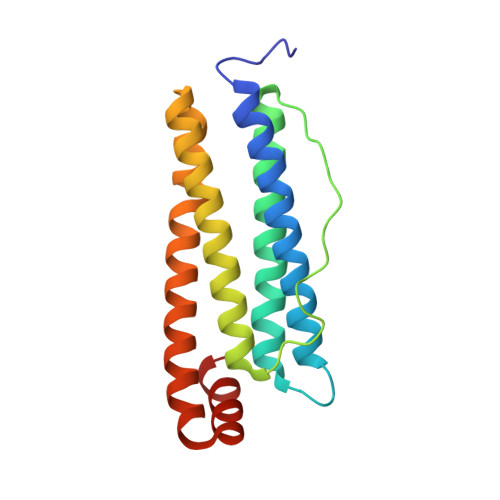
Arsenoplatin-Ferritin Nanocage: Structure and Cytotoxicity.
Ferraro, G., Pratesi, A., Cirri, D., Imbimbo, P., Maria Monti, D., Messori, L., Merlino, A.(2021) Int J Mol Sci 22
- PubMed: 33668605 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms22041874
- Primary Citation Related Structures:
7BD7 - PubMed Abstract:
Arsenoplatin-1 (AP-1), the prototype of a novel class of metallodrugs containing a PtAs(OH) 2 core, was encapsulated within the apoferritin (AFt) nanocage. UV-Vis absorption spectroscopy and inductively coupled plasma-atomic emission spectroscopy measurements confirmed metallodrug encapsulation and allowed us to determine the average amount of AP-1 trapped inside the cage. The X-ray structure of AP-1-encapsulated AFt was solved at 1.50 Å. Diffraction data revealed that an AP-1 fragment coordinates the side chain of a His residue. The biological activity of AP-1-loaded AFt was comparatively tested on a few representative cancer and non-cancer cell lines. Even though the presence of the cage reduces the overall cytotoxicity of AP-1, it improves its selectivity towards cancer cells.
- Department of Chemistry "Ugo Schiff", University of Florence, Via Della Lastruccia, 3-13, Sesto Fiorentino, 50019 Florence, Italy.
Organizational Affiliation: