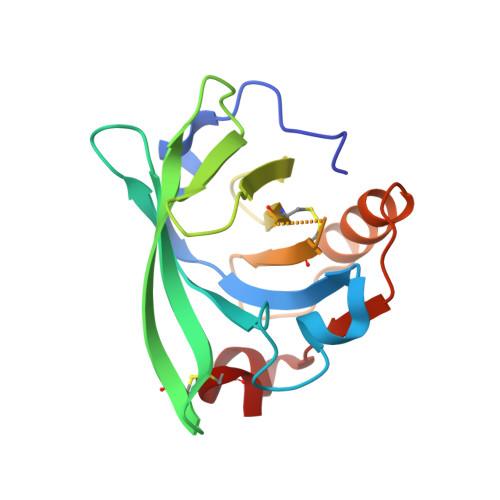
Interactions of new lactoglobulin variants with tetracaine: crystallographic studies of ligand binding to lactoglobulin mutants possessing single substitution in the binding pocket.
Loch, J., Bonarek, P., Siuda, M., Wrobel, P., Lewinski, K.(2021) Acta Biochim Pol 68: 23-28
- PubMed: 33719368 Search on PubMed
- DOI: https://doi.org/10.18388/abp.2020_5593
- Primary Citation Related Structures:
7BF7, 7BF8, 7BF9, 7BGA, 7BGX, 7BGZ, 7BH0 - PubMed Abstract:
β-Lactoglobulin (BLG) like other lipocalins can be modified by mutagenesis to re-direct its ligand binding properties. Local site-directed mutagenesis was used to change the geometry of the BLG ligand binding pocket and therefore change BLG ligand preferences. The presented studies are focused on previously described mutants L39Y, I56F, L58F, F105L, and M107L and two new BLG variants, L39K and F105A, and their interactions with local anesthetic drug tetracaine. Binding of tetracaine to BLG mutants was investigated by X-ray crystallography. Structural analysis revealed that for tetracaine binding, the shape of the binding pocket seems to be a more important factor than the substitutions influencing the number of interactions. Analyzed BLG mutants can be classified according to their binding properties to variants: capable of binding tetracaine in the β-barrel (L58F, M107L); capable of accommodating tetracaine on the protein surface (I56F) and unable to bind tetracaine (F105L). Variants L39K, L39Y, and F105A, had a binding pocket blocked by endogenous fatty acids. The new tetracaine binding site was found in the I56F variant. The site localized on the surface near Arg124 and Trp19 was previously predicted by in silico studies and was confirmed in the crystal structure.
- Department of Crystal Chemistry and Crystal Physics, Faculty of Chemistry, Jagiellonian University, Kraków, Poland.
Organizational Affiliation: