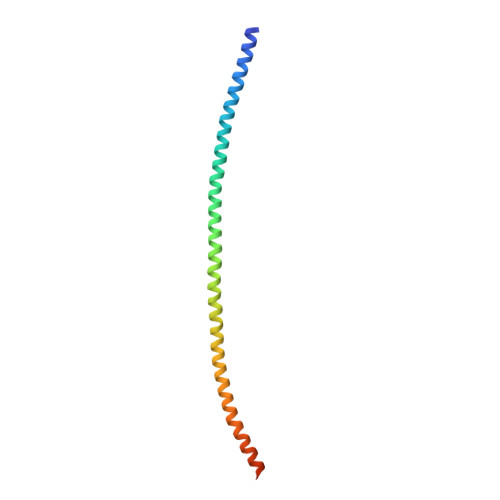Structural basis for the coiled-coil architecture of human CtIP.
Morton, C.R., Rzechorzek, N.J., Maman, J.D., Kuramochi, M., Sekiguchi, H., Rambo, R., Sasaki, Y.C., Davies, O.R., Pellegrini, L.(2021) Open Biol 11: 210060-210060
- PubMed: 34129781 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1098/rsob.210060
- Primary Citation Related Structures:
7BGF - PubMed Abstract:
The DNA repair factor CtIP has a critical function in double-strand break (DSB) repair by homologous recombination, promoting the assembly of the repair apparatus at DNA ends and participating in DNA-end resection. However, the molecular mechanisms of CtIP function in DSB repair remain unclear. Here, we present an atomic model for the three-dimensional architecture of human CtIP, derived from a multi-disciplinary approach that includes X-ray crystallography, small-angle X-ray scattering (SAXS) and diffracted X-ray tracking (DXT). Our data show that CtIP adopts an extended dimer-of-dimers structure, in agreement with a role in bridging distant sites on chromosomal DNA during the recombinational repair. The zinc-binding motif in the CtIP N-terminus alters dynamically the coiled-coil structure, with functional implications for the long-range interactions of CtIP with DNA. Our results provide a structural basis for the three-dimensional arrangement of chains in the CtIP tetramer, a key aspect of CtIP function in DNA DSB repair.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, UK.
Organizational Affiliation: