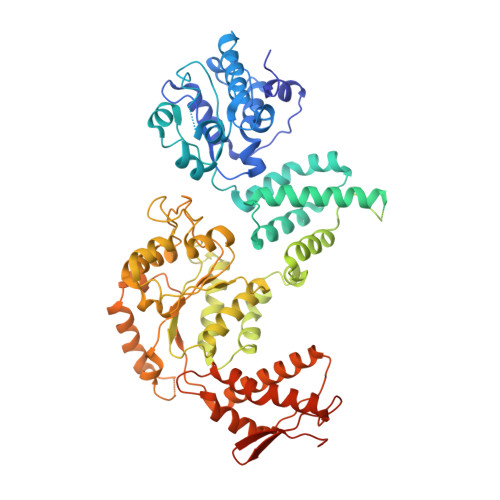
Split conformation of Chaetomium thermophilum Hsp104 disaggregase.
Inoue, Y., Hanazono, Y., Noi, K., Kawamoto, A., Kimatsuka, M., Harada, R., Takeda, K., Kita, R., Iwamasa, N., Shibata, K., Noguchi, K., Shigeta, Y., Namba, K., Ogura, T., Miki, K., Shinohara, K., Yohda, M.(2021) Structure 29: 721-730.e6
- PubMed: 33651974 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2021.02.002
- Primary Citation Related Structures:
5ZUI, 7CG3 - PubMed Abstract:
Hsp104 and its bacterial homolog ClpB form hexameric ring structures and mediate protein disaggregation. The disaggregated polypeptide is thought to thread through the central channel of the ring. However, the dynamic behavior of Hsp104 during disaggregation remains unclear. Here, we reported the stochastic conformational dynamics and a split conformation of Hsp104 disaggregase from Chaetomium thermophilum (CtHsp104) in the presence of ADP by X-ray crystallography, cryo-electron microscopy (EM), and high-speed atomic force microscopy (AFM). ADP-bound CtHsp104 assembles into a 6 5 left-handed spiral filament in the crystal structure at a resolution of 2.7 Å. The unit of the filament is a hexamer of the split spiral structure. In the cryo-EM images, staggered and split hexameric rings were observed. Further, high-speed AFM observations showed that a substrate addition enhanced the conformational change and increased the split structure's frequency. Our data suggest that split conformation is an off-pathway state of CtHsp104 during disaggregation.
- Department of Biotechnology and Life Science, Tokyo University of Agriculture and Technology, Koganei, Tokyo 184-8588, Japan.
Organizational Affiliation: