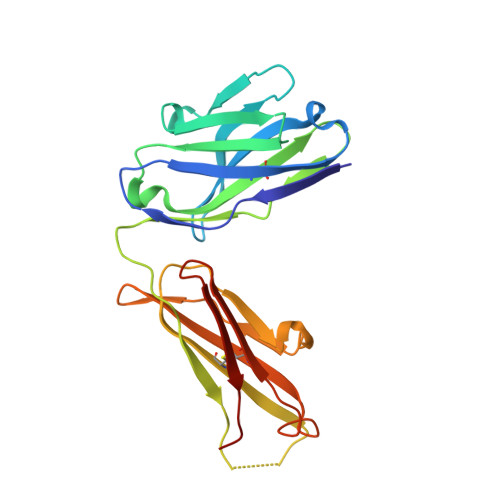
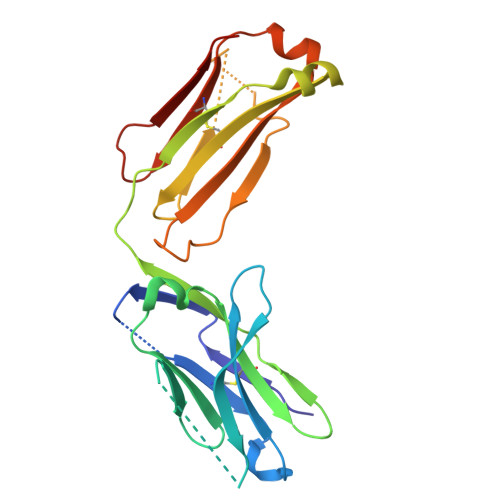
Conformational adaptability determining antibody recognition to distomer: structure analysis of enantioselective antibody against chiral drug gatifloxacin
Wang, L.T., Xie, W., Jiao, W.Y., Zhang, C., Li, X., Xu, Z., Huang, X., Lei, H.T., Shen, X.(2021) RSC Adv 11: 39534-39544