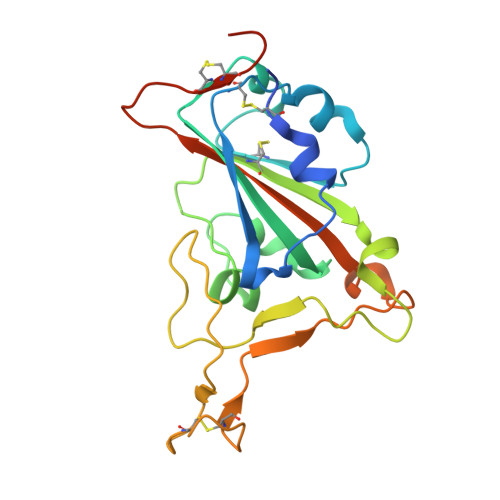
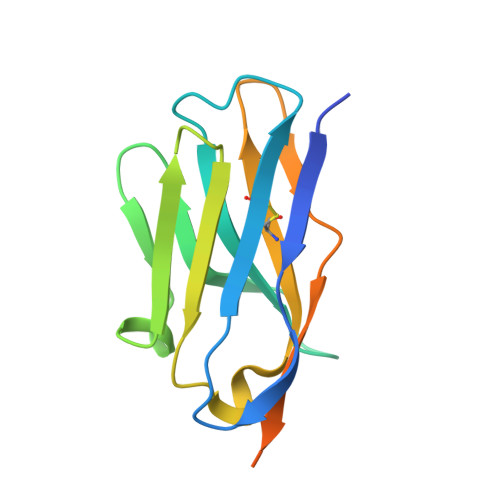
Structural Characterization of a Neutralizing Nanobody With Broad Activity Against SARS-CoV-2 Variants.
Li, T., Zhou, B., Luo, Z., Lai, Y., Huang, S., Zhou, Y., Li, Y., Gautam, A., Bourgeau, S., Wang, S., Bao, J., Tan, J., Lavillette, D., Li, D.(2022) Front Microbiol 13: 875840-875840
- PubMed: 35722331
- DOI: https://doi.org/10.3389/fmicb.2022.875840
- Primary Citation of Related Structures:
7F5H - PubMed Abstract:
SARS-CoV-2 and its variants, such as the Omicron continue to threaten public health. The virus recognizes the host cell by attaching its Spike (S) receptor-binding domain (RBD) to the host receptor, ACE2. Therefore, RBD is a primary target for neutralizing antibodies and vaccines. Here, we report the isolation and biological and structural characterization of a single-chain antibody (nanobody) from RBD-immunized alpaca. The nanobody, named DL28, binds to RBD tightly with a K D of 1.56 nM and neutralizes the original SARS-CoV-2 strain with an IC 50 of 0.41 μg mL -1 . Neutralization assays with a panel of variants of concern (VOCs) reveal its wide-spectrum activity with IC 50 values ranging from 0.35 to 1.66 μg mL -1 for the Alpha/Beta/Gamma/Delta and an IC 50 of 0.66 μg mL -1 for the currently prevalent Omicron. Competition binding assays show that DL28 blocks ACE2-binding. However, structural characterizations and mutagenesis suggest that unlike most antibodies, the blockage by DL28 does not involve direct competition or steric hindrance. Rather, DL28 may use a "conformation competition" mechanism where it excludes ACE2 by keeping an RBD loop in a conformation incompatible with ACE2-binding.
- State Key Laboratory of Molecular Biology, Chinese Academy of Sciences Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: