The Role of the HLA Class I alpha 2 Helix in Determining Ligand Hierarchy for the Killer Cell Ig-like Receptor 3DL1.
Saunders, P.M., MacLachlan, B.J., Widjaja, J., Wong, S.C., Oates, C.V.L., Rossjohn, J., Vivian, J.P., Brooks, A.G.(2021) J Immunol 206: 849-860
- PubMed: 33441440 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.2001109
- Primary Citation Related Structures:
7K80, 7K81 - PubMed Abstract:
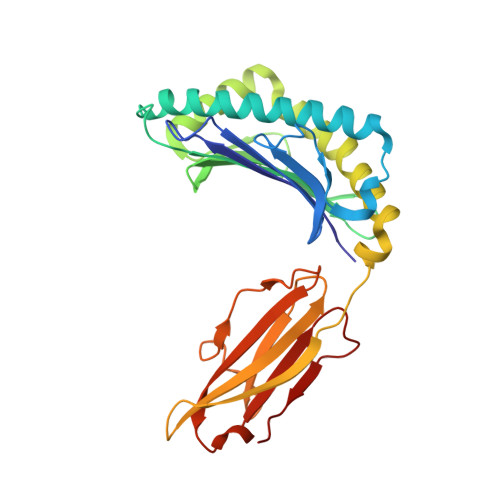
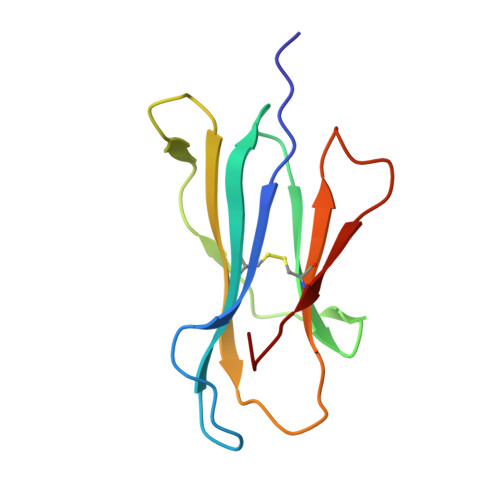
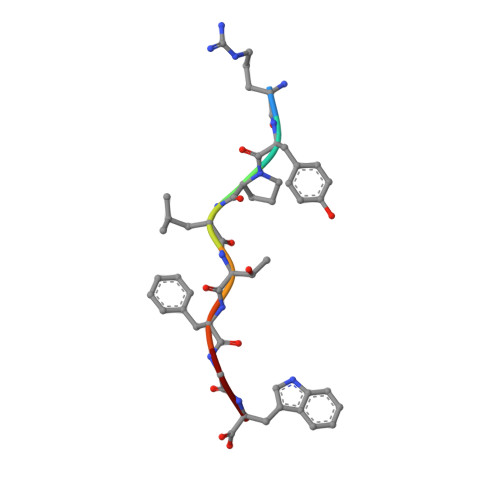
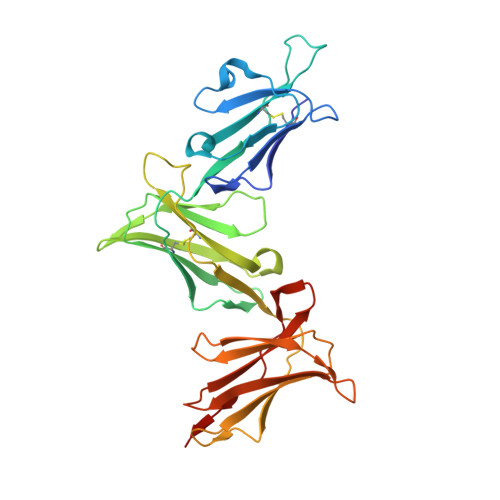
HLA class I molecules that represent ligands for the inhibitory killer cell Ig-like receptor (KIR) 3DL1 found on NK cells are categorically defined as those HLA-A and HLA-B allotypes containing the Bw4 motif, yet KIR3DL1 demonstrates hierarchical recognition of these HLA-Bw4 ligands. To better understand the molecular basis underpinning differential KIR3DL1 recognition, the HLA-A Bw4 family of allotypes were investigated. Transfected human 721.221 cells expressing HLA-A*32:01 strongly inhibited primary human KIR3DL1 + NK cells, whereas HLA-A*24:02 and HLA-A*23:01 displayed intermediate potency and HLA-A*25:01 failed to inhibit activation of KIR3DL1 + NK cells. Structural studies demonstrated that recognition of HLA-A*24:02 by KIR3DL1 used identical contacts as the potent HLA-B*57:01 ligand. Namely, the D1-D2 domains of KIR3DL1 were placed over the α1 helix and α2 helix of the HLA-A*24:02 binding cleft, respectively, whereas the D0 domain contacted the side of the HLA-A*24:02 molecule. Nevertheless, functional analyses showed KIR3DL1 recognition of HLA-A*24:02 was more sensitive to substitutions within the α2 helix of HLA-A*24:02, including residues Ile 142 and Lys 144 Furthermore, the presence of Thr 149 in the α2 helix of HLA-A*25:01 abrogated KIR3DL1 + NK inhibition. Together, these data demonstrate a role for the HLA class I α2 helix in determining the hierarchy of KIR3DL1 ligands. Thus, recognition of HLA class I is dependent on a complex interplay between the peptide repertoire, polymorphisms within and proximal to the Bw4 motif, and the α2 helix. Collectively, the data furthers our understanding of KIR3DL1 ligands and will inform genetic association and immunogenetics studies examining the role of KIR3DL1 in disease settings.
- Department of Microbiology and Immunology, Peter Doherty Institute for Infection and Immunity, The University of Melbourne, Melbourne, Victoria 3000, Australia; philippa.saunders@unimelb.edu.au agbrooks@unimelb.edu.au.
Organizational Affiliation: