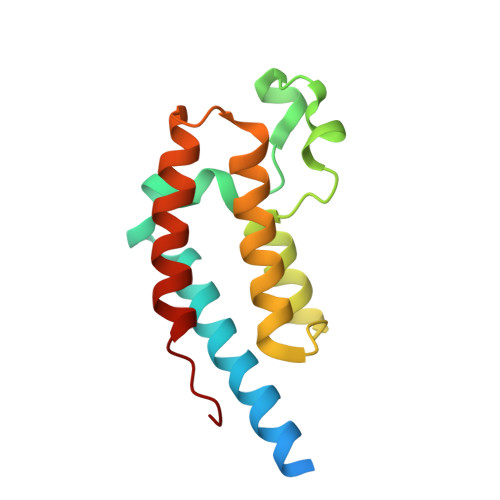
Discovery and characterization of bromodomain 2-specific inhibitors of BRDT.
Yu, Z., Ku, A.F., Anglin, J.L., Sharma, R., Ucisik, M.N., Faver, J.C., Li, F., Nyshadham, P., Simmons, N., Sharma, K.L., Nagarajan, S., Riehle, K., Kaur, G., Sankaran, B., Storl-Desmond, M., Palmer, S.S., Young, D.W., Kim, C., Matzuk, M.M.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33637650
- DOI: https://doi.org/10.1073/pnas.2021102118
- Primary Citation of Related Structures:
7L99, 7L9A - PubMed Abstract:
Bromodomain testis (BRDT), a member of the bromodomain and extraterminal (BET) subfamily that includes the cancer targets BRD2, BRD3, and BRD4, is a validated contraceptive target. All BET subfamily members have two tandem bromodomains (BD1 and BD2). Knockout mice lacking BRDT-BD1 or both bromodomains are infertile. Treatment of mice with JQ1, a BET BD1/BD2 nonselective inhibitor with the highest affinity for BRD4, disrupts spermatogenesis and reduces sperm number and motility. To assess the contribution of each BRDT bromodomain, we screened our collection of DNA-encoded chemical libraries for BRDT-BD1 and BRDT-BD2 binders. High-enrichment hits were identified and resynthesized off-DNA and examined for their ability to compete with JQ1 in BRDT and BRD4 bromodomain AlphaScreen assays. These studies identified CDD-1102 as a selective BRDT-BD2 inhibitor with low nanomolar potency and >1,000-fold selectivity over BRDT-BD1. Structure-activity relationship studies of CDD-1102 produced a series of additional BRDT-BD2/BRD4-BD2 selective inhibitors, including CDD-1302, a truncated analog of CDD-1102 with similar activity, and CDD-1349, an analog with sixfold selectivity for BRDT-BD2 versus BRD4-BD2. BROMOscan bromodomain profiling confirmed the great affinity and selectivity of CDD-1102 and CDD-1302 on all BET BD2 versus BD1 with the highest affinity for BRDT-BD2. Cocrystals of BRDT-BD2 with CDD-1102 and CDD-1302 were determined at 2.27 and 1.90 Å resolution, respectively, and revealed BRDT-BD2 specific contacts that explain the high affinity and selectivity of these compounds. These BD2-specific compounds and their binding to BRDT-BD2 are unique compared with recent reports and enable further evaluation of their nonhormonal contraceptive potential in vitro and in vivo.
- Center for Drug Discovery, Department of Pathology & Immunology, Baylor College of Medicine, Houston, TX 77030.
Organizational Affiliation: