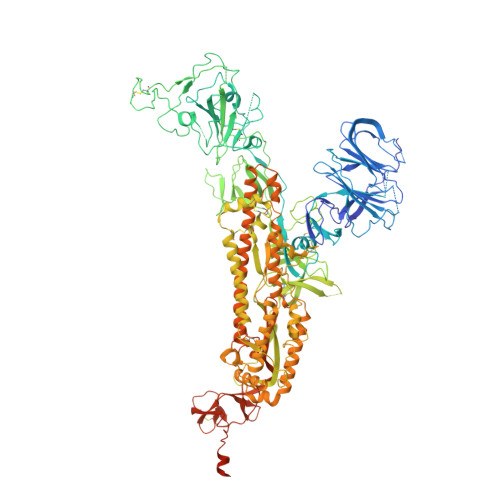
Structures of the Omicron spike trimer with ACE2 and an anti-Omicron antibody.
Yin, W., Xu, Y., Xu, P., Cao, X., Wu, C., Gu, C., He, X., Wang, X., Huang, S., Yuan, Q., Wu, K., Hu, W., Huang, Z., Liu, J., Wang, Z., Jia, F., Xia, K., Liu, P., Wang, X., Song, B., Zheng, J., Jiang, H., Cheng, X., Jiang, Y., Deng, S.J., Xu, H.E.(2022) Science 375: 1048-1053
- PubMed: 35133176 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abn8863
- Primary Citation Related Structures:
7WP9, 7WPA, 7WPB, 7WPC, 7WPD, 7WPE, 7WPF, 7WRV - PubMed Abstract:
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron variant has become the dominant infective strain. We report the structures of the Omicron spike trimer on its own and in complex with angiotensin-converting enzyme 2 (ACE2) or an anti-Omicron antibody. Most Omicron mutations are located on the surface of the spike protein and change binding epitopes to many current antibodies. In the ACE2-binding site, compensating mutations strengthen receptor binding domain (RBD) binding to ACE2. Both the RBD and the apo form of the Omicron spike trimer are thermodynamically unstable. An unusual RBD-RBD interaction in the ACE2-spike complex supports the open conformation and further reinforces ACE2 binding to the spike trimer. A broad-spectrum therapeutic antibody, JMB2002, which has completed a phase 1 clinical trial, maintains neutralizing activity against Omicron. JMB2002 binds to RBD differently from other characterized antibodies and inhibits ACE2 binding.
- The CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China.
Organizational Affiliation: