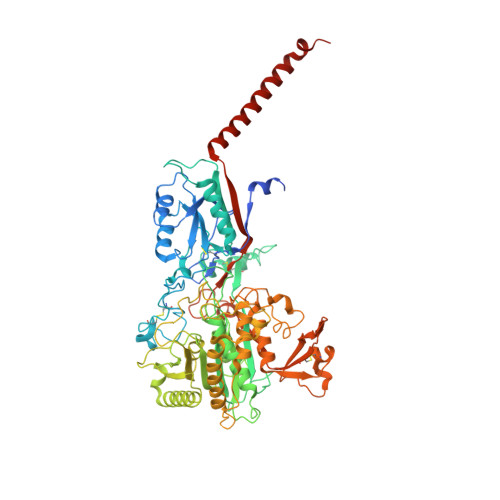
Molecular basis for isoform-selective inhibition of presenilin-1 by MRK-560.
Guo, X., Wang, Y., Zhou, J., Jin, C., Wang, J., Jia, B., Jing, D., Yan, C., Lei, J., Zhou, R., Shi, Y.(2022) Nat Commun 13: 6299-6299
- PubMed: 36272978 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33817-5
- Primary Citation Related Structures:
7Y5T, 7Y5X, 7Y5Z - PubMed Abstract:
Inhibition of γ-secretase activity represents a potential therapeutic strategy for Alzheimer's disease (AD). MRK-560 is a selective inhibitor with higher potency for Presenilin 1 (PS1) than for PS2, the two isoforms of the catalytic subunit of γ-secretase, although the underlying mechanism remains elusive. Here we report the cryo-electron microscopy (cryo-EM) structures of PS1 and PS2-containing γ-secretase complexes with and without MRK-560 at overall resolutions of 2.9-3.4 Å. MRK-560 occupies the substrate binding site of PS1, but is invisible in PS2. Structural comparison identifies Thr281 and Leu282 in PS1 to be the determinant for isoform-dependent sensitivity to MRK-560, which is confirmed by swapping experiment between PS1 and PS2. By revealing the mechanism for isoform-selective inhibition of presenilin, our work may facilitate future drug discovery targeting γ-secretase.
- Beijing Frontier Research Center for Biological Structure, Tsinghua-Peking Joint Center for Life Sciences, Key Laboratory for Protein Sciences of Ministry of Education, School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: