Histidine-51 facilitates deprotonation of the zinc-bound ligand during catalysis by horse liver alcohol dehydrogenase
Plapp, B.V., Kovaleva, E.G.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Alcohol dehydrogenase E chain | 374 | Equus caballus | Mutation(s): 1 EC: 1.1.1.1 | 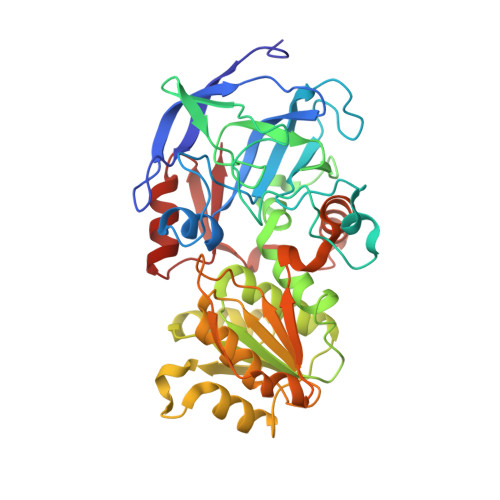 | |
UniProt | |||||
Find proteins for P00327 (Equus caballus) Explore P00327 Go to UniProtKB: P00327 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00327 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAJ (Subject of Investigation/LOI) Query on NAJ | E [auth A], L [auth B] | NICOTINAMIDE-ADENINE-DINUCLEOTIDE (ACIDIC FORM) C21 H27 N7 O14 P2 BAWFJGJZGIEFAR-NNYOXOHSSA-N |  | ||
| DKA (Subject of Investigation/LOI) Query on DKA | F [auth A], M [auth B] | DECANOIC ACID C10 H20 O2 GHVNFZFCNZKVNT-UHFFFAOYSA-N |  | ||
| MRD Query on MRD | I [auth A] | (4R)-2-METHYLPENTANE-2,4-DIOL C6 H14 O2 SVTBMSDMJJWYQN-RXMQYKEDSA-N |  | ||
| MPD Query on MPD | G [auth A], H [auth A] | (4S)-2-METHYL-2,4-PENTANEDIOL C6 H14 O2 SVTBMSDMJJWYQN-YFKPBYRVSA-N |  | ||
| ZN Query on ZN | C [auth A], D [auth A], J [auth B], K [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 44.24 | α = 92.01 |
| b = 51.25 | β = 102.87 |
| c = 92.46 | γ = 109.91 |
| Software Name | Purpose |
|---|---|
| d*TREK | data scaling |
| REFMAC | refinement |
| d*TREK | data reduction |
| REFMAC | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute on Alcohol Abuse and Alcoholism (NIH/NIAAA) | United States | AA00279 |