7.10 MAG. A Novel Host Monoacylglyceride for In Meso (Lipid Cubic Phase) Crystallization of Membrane Proteins.
Krawinski, P., Smithers, L., van Dalsen, L., Boland, C., Ostrovitsa, N., Perez, J., Caffrey, M.(2024) Cryst Growth Des 24: 2985-3001
- PubMed: 38585376
- DOI: https://doi.org/10.1021/acs.cgd.4c00087
- Primary Citation Related Structures:
8RQP, 8RQQ, 8RQR - PubMed Abstract:
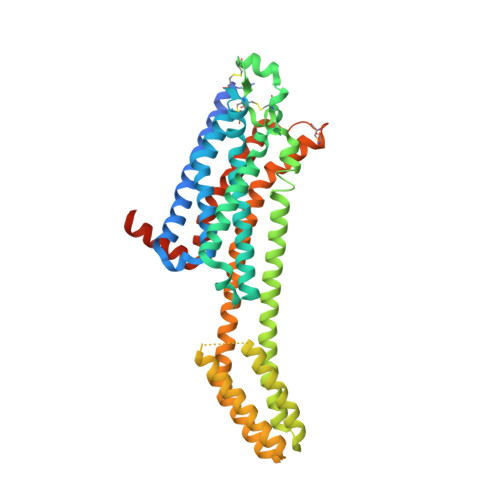
A novel monoacylglycerol, 7.10 MAG, has been produced for use in the in meso (lipid cubic phase) crystallization of membrane proteins and complexes. 7.10 MAG differs from monoolein, the most extensively used lipid for in meso crystallization, in that it is shorter in chain length by one methylene and its cis olefinic bond is two carbons closer to the glycerol headgroup. These changes in structure alter the phase behavior of the hydrated lipid and the microstructure of the corresponding mesophases formed. Temperature-composition phase diagrams for 7.10 MAG have been constructed using small- and wide-angle X-ray scattering over a range of temperatures and hydration levels that span those used for crystallization. The phase diagrams include lamellar crystalline, fluid isotropic, lamellar liquid-crystalline, cubic- Ia 3 d, and cubic- Pn 3 m phases, as observed with monoolein. Conspicuous by its absence is the inverted hexagonal phase which is rationalized on the basis of 7.10 MAG's chemical constitution. The cubic phase prepared with the new lipid facilitates the growth of crystals that were used to generate high-resolution structures of intramembrane β-barrel and α-helical proteins. Compatibility of fully hydrated 7.10 MAG with cholesterol and phosphatidylcholine means that these two lipids can be used as additives to optimize crystallogenesis in screening trials with 7.10 MAG as the host lipid.
- Membrane Structural and Functional Biology Group, School of Medicine and School of Biochemistry and Immunology, Trinity College Dublin, Dublin D02 R590, Ireland.
Organizational Affiliation: