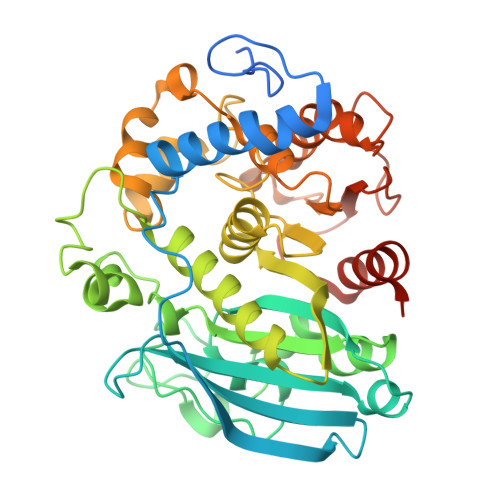
Interaction of a glucuronoyl esterase with complex fragments of the plant cell wall hemicellulose.
Zaghini, A., Ostberg, E.B., Banerjee, S., Mazurkewich, S., Yu, L., Dupree, P., Larsbrink, J., Leggio, L.L.(2026) Int J Biol Macromol : 151020-151020
- PubMed: 41720372
- DOI: https://doi.org/10.1016/j.ijbiomac.2026.151020
- Primary Citation of Related Structures:
9HUC, 9HVR - PubMed Abstract:
Glucuronoyl esterases (GEs) catalyze the cleavage of ester linkages between lignin and glucuronic acid moieties on glucuronoxylan in plant cell walls and are promising biochemical tools for industrial processing of these recalcitrant natural resources. However, details on how GEs interact with and catalyze degradation of their natural substrates are sparse. Using well-diffracting crystals of the model GE OtCE15A, we sought to structurally elucidate its interactions with fragments and analogues of the lignin carbohydrate complex, including commercially available oligosaccharides, digests of complex polysaccharides, and chemical fragments representing building blocks of lignin. While most compounds failed to bind in crystals, the analysis uncovered the structure of a complex with a heptasaccharide (a decorated xylopentaose), the largest ligand bound to a GE experimental structure to date. Most hydrogen bonding interactions are with the glucuronic acid moiety, which is almost totally buried on binding, however, the rest of the saccharide chain also has a significant contact surface. The structure shows for the first time that OtCE15A can accommodate an arabinose decoration on the glucuronoxylan chain it attacks. Structural comparison suggests that the polysaccharide conformation may influence specificity, reinforcing the view of GEs as true carbohydrate-active enzymes rather than opportunistic promiscuous esterases.
- Department of Chemistry, University of Copenhagen, DK-2100, Copenhagen, Denmark.
Organizational Affiliation: