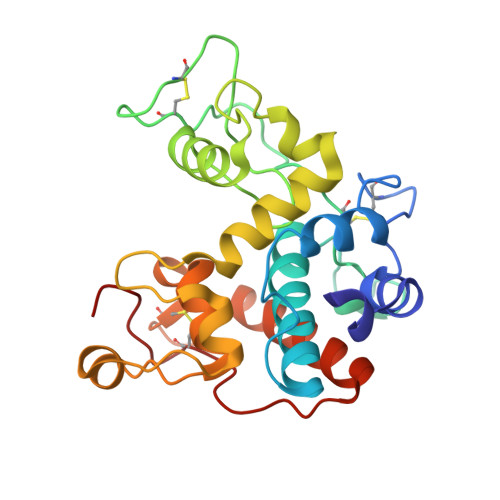
Structure of Jack Bean Chitinase
Hahn, M., Hennig, M., Schlesier, B., Hohne, W.(2000) Acta Crystallogr D Biol Crystallogr 56: 1096
- PubMed: 10957628 Search on PubMed
- DOI: https://doi.org/10.1107/s090744490000857x
- Primary Citation Related Structures:
1DXJ - PubMed Abstract:
The structure of jack bean chitinase was solved at 1.8 A resolution by molecular replacement. It is an alpha-helical protein with three disulfide bridges. The active site is related in structure to animal and viral lysozymes. However, unlike in lysozyme, the architecture of the active site suggests a single-step cleavage. According to this mechanism, Glu68 is the proton donor and Glu90 assists in the reaction by moving towards the substrate and recruiting a water molecule that acts as the nucleophile. In this model, a water molecule was found in contact with Glu90 O(epsilon1) and Thr119 O(gamma) at a distance of 3.0 and 2.8 A, respectively. The model is in accordance with the observed inversion mechanism.
- Institut für Biochemie, Charité, Humboldt-Universität, Monbijoustrasse 2, 10117 Berlin, Germany. michael.hahn@charite.de
Organizational Affiliation: