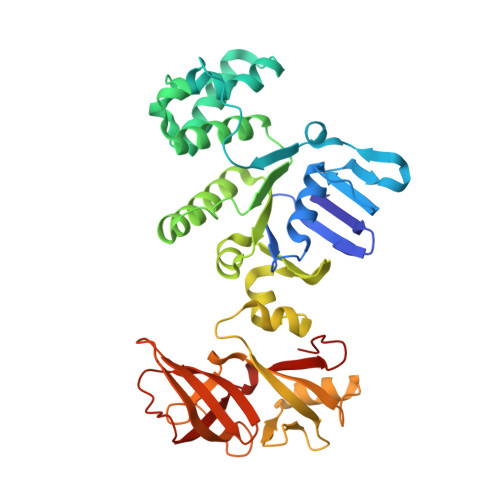
Crystal structure of MalK, the ATPase subunit of the trehalose/maltose ABC transporter of the archaeon Thermococcus litoralis.
Diederichs, K., Diez, J., Greller, G., Muller, C., Breed, J., Schnell, C., Vonrhein, C., Boos, W., Welte, W.(2000) EMBO J 19: 5951-5961
- PubMed: 11080142 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/19.22.5951
- Primary Citation Related Structures:
1G29 - PubMed Abstract:
The members of the ABC transporter family transport a wide variety of molecules into or out of cells and cellular compartments. Apart from a translocation pore, each member possesses two similar nucleoside triphosphate-binding subunits or domains in order to couple the energy-providing reaction with transport. In the maltose transporter of several Gram-negative bacteria and the archaeon Thermo coccus litoralis, the nucleoside triphosphate-binding subunit contains a C-terminal regulatory domain. A dimer of the subunit is attached cytoplasmically to the translocation pore. Here we report the crystal structure of this dimer showing two bound pyrophosphate molecules at 1.9 A resolution. The dimer forms by association of the ATPase domains, with the two regulatory domains attached at opposite poles. Significant deviation from 2-fold symmetry is seen at the interface of the dimer and in the regions corresponding to those residues known to be in contact with the translocation pore. The structure and its relationship to function are discussed in the light of known mutations from the homologous Escherichia coli and Salmonella typhimurium proteins.
- Fachbereich Biologie, Universität Konstanz, M656, D-78457 Konstanz, Germany.
Organizational Affiliation: