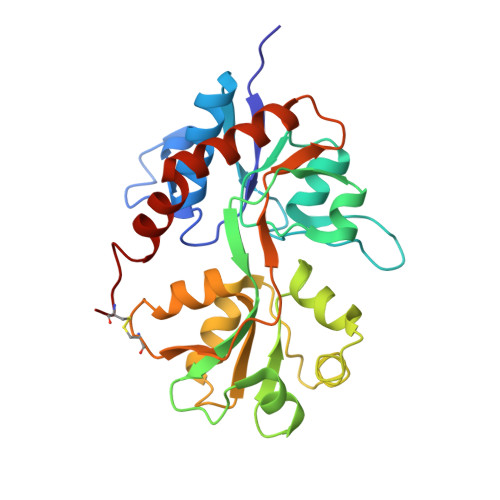
Mechanism of glutamate receptor desensitization.
Sun, Y., Olson, R., Horning, M., Armstrong, N., Mayer, M., Gouaux, E.(2002) Nature 417: 245-253
- PubMed: 12015593 Search on PubMed
- DOI: https://doi.org/10.1038/417245a
- Primary Citation Related Structures:
1LB8, 1LB9, 1LBB, 1LBC - PubMed Abstract:
Ligand-gated ion channels transduce chemical signals into electrical impulses by opening a transmembrane pore in response to binding one or more neurotransmitter molecules. After activation, many ligand-gated ion channels enter a desensitized state in which the neurotransmitter remains bound but the ion channel is closed. Although receptor desensitization is crucial to the functioning of many ligand-gated ion channels in vivo, the molecular basis of this important process has until now defied analysis. Using the GluR2 AMPA-sensitive glutamate receptor, we show here that the ligand-binding cores form dimers and that stabilization of the intradimer interface by either mutations or allosteric modulators reduces desensitization. Perturbations that destabilize the interface enhance desensitization. Receptor activation involves conformational changes within each subunit that result in an increase in the separation of portions of the receptor that are linked to the ion channel. Our analysis defines the dimer interface in the resting and activated state, indicates how ligand binding is coupled to gating, and suggests modes of dimer dimer interaction in the assembled tetramer. Desensitization occurs through rearrangement of the dimer interface, which disengages the agonist-induced conformational change in the ligand-binding core from the ion channel gate.
- Department of Biochemistry and Molecular Biophysics, Columbia University, 650 West 168th Street, New York, New York 10032, USA.
Organizational Affiliation: