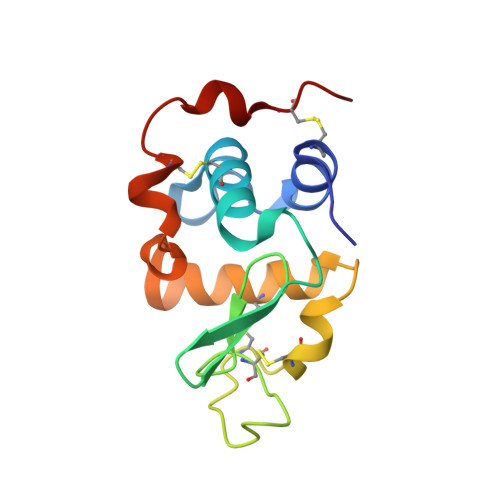
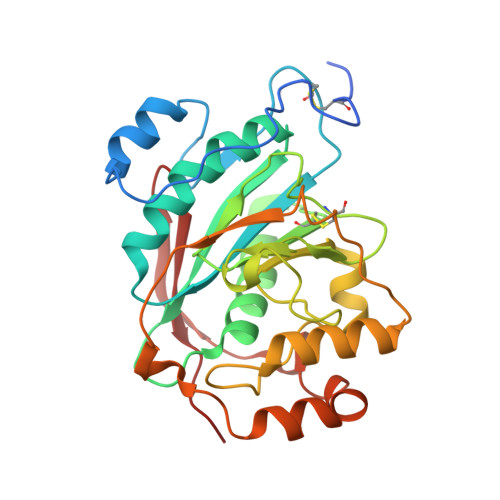
Crystal Structure of Lactose Synthase Reveals a Large Conformational Change in its Catalytic Component, the beta-1,4-galactosyltransferase
Ramakrishnan, B., Qasba, P.K.(2001) J Mol Biology 310: 205-218
- PubMed: 11419947 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4757
- Primary Citation Related Structures:
1NF5, 1NHE, 1NKH, 1NQI - PubMed Abstract:
The lactose synthase (LS) enzyme is a 1:1 complex of a catalytic component, beta1,4-galactosyltransferse (beta4Gal-T1) and a regulatory component, alpha-lactalbumin (LA), a mammary gland-specific protein. LA promotes the binding of glucose (Glc) to beta4Gal-T1, thereby altering its sugar acceptor specificity from N-acetylglucosamine (GlcNAc) to glucose, which enables LS to synthesize lactose, the major carbohydrate component of milk. The crystal structures of LS bound with various substrates were solved at 2 A resolution. These structures reveal that upon substrate binding to beta4Gal-T1, a large conformational change occurs in the region comprising residues 345 to 365. This repositions His347 in such a way that it can participate in the coordination of a metal ion, and creates a sugar and LA-binding site. At the sugar-acceptor binding site, a hydrophobic N-acetyl group-binding pocket is found, formed by residues Arg359, Phe360 and Ile363. In the Glc-bound structure, this hydrophobic pocket is absent. For the binding of Glc to LS, a reorientation of the Arg359 side-chain occurs, which blocks the hydrophobic pocket and maximizes the interactions with the Glc molecule. Thus, the role of LA is to hold Glc by hydrogen bonding with the O-1 hydroxyl group in the acceptor-binding site on beta4Gal-T1, while the N-acetyl group-binding pocket in beta4Gal-T1 adjusts to maximize the interactions with the Glc molecule. This study provides details of a structural basis for the partially ordered kinetic mechanism proposed for lactose synthase.
- Structural Glycobiology Section, Intramural Research Support Program-SAIC, Laboratory of Experimental and Computational Biology, CCR, NCI, Frederick, MD 21702, USA.
Organizational Affiliation: