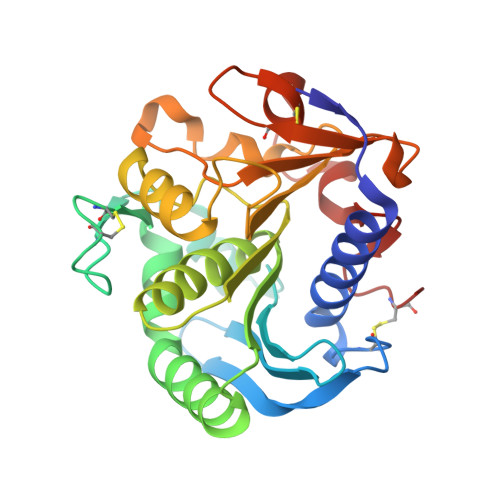
Structure of a Feruloyl Esterase from Aspergillus Niger
Mcauley, K.E., Svendsen, A., Patkar, S.A., Wilson, K.S.(2004) Acta Crystallogr D Biol Crystallogr 60: 878
- PubMed: 15103133
- DOI: https://doi.org/10.1107/S0907444904004937
- Primary Citation of Related Structures:
1UWC, 1UZA - PubMed Abstract:
The crystallographic structure of feruloyl esterase from Aspergillus niger has been determined to a resolution of 1.5 A by molecular replacement. The protein has an alpha/beta-hydrolase structure with a Ser-His-Asp catalytic triad; the overall fold of the protein is very similar to that of the fungal lipases. The structure of the enzyme-product complex was determined to a resolution of 1.08 A and reveals dual conformations for the serine and histidine residues at the active site.
- CCLRC Daresbury Laboratory, Warrington, Cheshire WA4 4AD, England.
Organizational Affiliation: