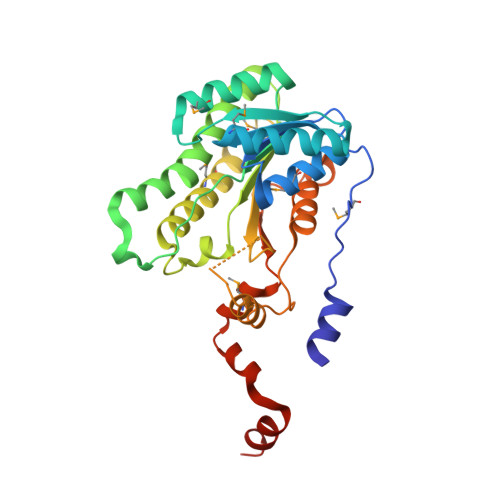
Structure and Reactivity of Human Mitochondrial 2,4-Dienoyl-Coa Reductase: Enzyme-Ligand Interactions in a Distinctive Short-Chain Reductase Active Site
Alphey, M.S., Yu, W., Byres, E., Li, D., Hunter, W.N.(2005) J Biological Chem 280: 3068
- PubMed: 15531764 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411069200
- Primary Citation Related Structures:
1W6U, 1W73, 1W8D - PubMed Abstract:
Fatty acid catabolism by beta-oxidation mainly occurs in mitochondria and to a lesser degree in peroxisomes. Poly-unsaturated fatty acids are problematic for beta-oxidation, because the enzymes directly involved are unable to process all the different double bond conformations and combinations that occur naturally. In mammals, three accessory proteins circumvent this problem by catalyzing specific isomerization and reduction reactions. Central to this process is the NADPH-dependent 2,4-dienoyl-CoA reductase. We present high resolution crystal structures of human mitochondrial 2,4-dienoyl-CoA reductase in binary complex with cofactor, and the ternary complex with NADP(+) and substrate trans-2,trans-4-dienoyl-CoA at 2.1 and 1.75 A resolution, respectively. The enzyme, a homotetramer, is a short-chain dehydrogenase/reductase with a distinctive catalytic center. Close structural similarity between the binary and ternary complexes suggests an absence of large conformational changes during binding and processing of substrate. The site of catalysis is relatively open and placed beside a flexible loop thereby allowing the enzyme to accommodate and process a wide range of fatty acids. Seven single mutants were constructed, by site-directed mutagenesis, to investigate the function of selected residues in the active site thought likely to either contribute to the architecture of the active site or to catalysis. The mutant proteins were overexpressed, purified to homogeneity, and then characterized. The structural and kinetic data are consistent and support a mechanism that derives one reducing equivalent from the cofactor, and one from solvent. Key to the acquisition of a solvent-derived proton is the orientation of substrate and stabilization of a dienolate intermediate by Tyr-199, Asn-148, and the oxidized nicotinamide.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee, DD1 5EH, United Kingdom.
Organizational Affiliation: