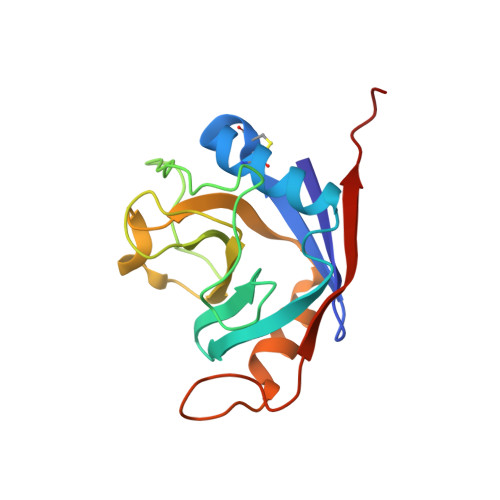
Structure of recombinant human cyclophilin J, a novel member of the cyclophilin family.
Huang, L.L., Zhao, X.M., Huang, C.Q., Yu, L., Xia, Z.X.(2005) Acta Crystallogr D Biol Crystallogr 61: 316-321
- PubMed: 15735342
- DOI: https://doi.org/10.1107/S0907444904033189
- Primary Citation Related Structures:
1XYH - PubMed Abstract:
Cyclophilins (CyPs) are a large class of highly conserved ubiquitous peptidyl-prolyl cis-trans isomerases. CyPs have also been identified as being a specific receptor for the immunosuppressive drug cyclosporin A and are involved in a variety of biological functions. CyPJ is a novel member of the CyP family and human CyPJ (hCyPJ) is the protein encoded by a cyclophilin-like gene from human foetal brain, which shows 50% sequence identity to human cyclophilin A (hCyPA). Recombinant hCyPJ was expressed in Escherichia coli and purified. The three-dimensional structure of hCyPJ has been determined by molecular replacement using the hCyPA structure as the search model and has been refined at 2.6 angstroms resolution. The hCyPJ molecule contains four helices and one beta-barrel composed of eight antiparallel beta-strands. The overall secondary and tertiary structures of hCyPJ are similar to those of hCyPA, but hCyPJ contains an additional disulfide bridge and four segments with conformations that are strikingly different from those of hCyPA. His43 and Gln52 of hCyPJ are expected to be the active sites based on sequence alignment with hCyPA. The hCyPJ structure shows a conserved water molecule close to His43 and Gln52 which appears to support the solvent-assisted mechanism.
- State Key Laboratory of Bio-organic and Natural Products Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai 200032, People's Republic of China.
Organizational Affiliation: