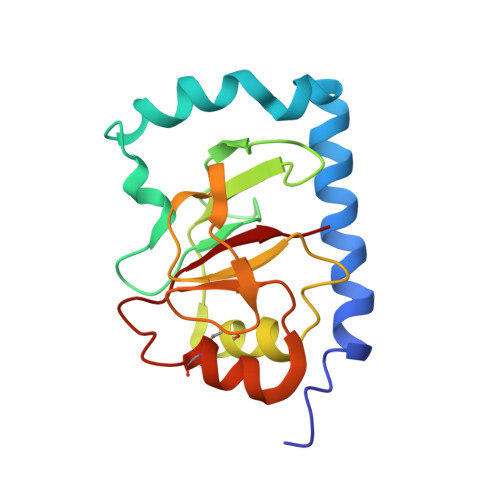
Structure of the Bundle-forming Pilus from Enteropathogenic Escherichia coli
Ramboarina, S., Fernandes, P.J., Daniell, S., Islam, S., Simpson, P., Frankel, G., Booy, F., Donnenberg, M.S., Matthews, S.(2005) J Biological Chem 280: 40252-40260
- PubMed: 16172128 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M508099200
- Primary Citation Related Structures:
1ZWT - PubMed Abstract:
Bundle-forming pili (BFP) are essential for the full virulence of enteropathogenic Escherichia coli (EPEC) because they are required for localized adherence to epithelial cells and auto-aggregation. We report the high resolution structure of bundlin, the monomer of BFP, solved by NMR. The structure reveals a new variation in the topology of type IVb pilins with significant differences in the composition and relative orientation of elements of secondary structure. In addition, the structural parameters of native BFP filaments were determined by electron microscopy after negative staining. The solution structure of bundlin was assembled according to these helical parameters to provide a plausible atomic resolution model for the BFP filament. We show that EPEC and Vibriocholerae type IVb pili display distinct differences in their monomer subunits consistent with data showing that bundlin and TcpA cannot complement each other, but assemble into filaments with similar helical organization.
- Department of Biological Sciences, Wolfson Laboratory, Imperial College, London SW72AZ, United Kingdom.
Organizational Affiliation: