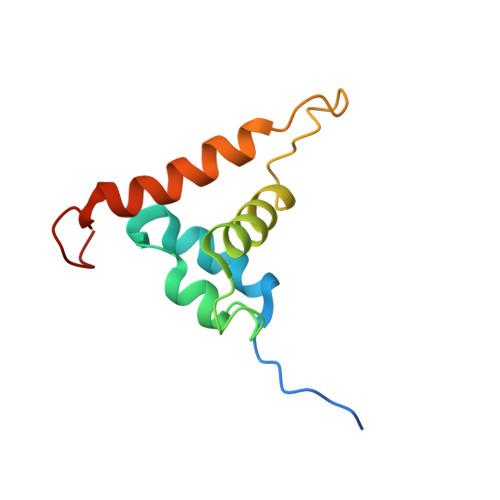
Solution structure of the Arabidopsis thaliana telomeric repeat-binding protein DNA binding domain: a new fold with an additional C-terminal helix
Sue, S.C., Hsiao, H.H., Chung, B.C., Cheng, Y.H., Hsueh, K.L., Chen, C.M., Ho, C.H., Huang, T.H.(2006) J Mol Biology 356: 72-85
- PubMed: 16337232 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.009
- Primary Citation Related Structures:
2AJE - PubMed Abstract:
The double-stranded telomeric repeat-binding protein (TRP) AtTRP1 is isolated from Arabidopsis thaliana. Using gel retardation assays, we defined the C-terminal 97 amino acid residues, Gln464 to Val560 (AtTRP1(464-560)), as the minimal structured telomeric repeat-binding domain. This region contains a typical Myb DNA-binding motif and a C-terminal extension of 40 amino acid residues. The monomeric AtTRP1(464-560) binds to a 13-mer DNA duplex containing a single repeat of an A.thaliana telomeric DNA sequence (GGTTTAG) in a 1:1 complex, with a K(D) approximately 10(-6)-10(-7) M. Nuclear magnetic resonance (NMR) examination revealed that the solution structure of AtTRP1(464-560) is a novel four-helix tetrahedron rather than the three-helix bundle structure found in typical Myb motifs and other TRPs. Binding of the 13-mer DNA duplex to AtTRP1(464-560) induced significant chemical shift perturbations of protein amide resonances, which suggests that helix 3 (H3) and the flexible loop connecting H3 and H4 are essential for telomeric DNA sequence recognition. Furthermore, similar to that in hTRF1, the N-terminal arm likely contributes to or stabilizes DNA binding. Sequence comparisons suggested that the four-helix structure and the involvement of the loop residues in DNA binding may be features unique to plant TRPs.
- Institute of Biomedical Sciences, Academia Sinica, Nankang, Taipei, Taiwan, ROC.
Organizational Affiliation: