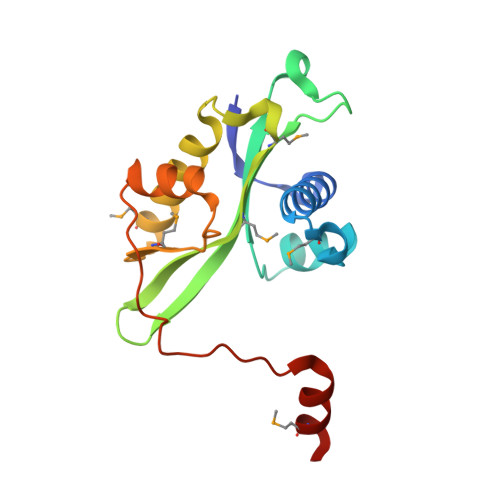
Structures of wild-type and mutant human spermidine/spermine N1-acetyltransferase, a potential therapeutic drug target.
Bewley, M.C., Graziano, V., Jiang, J., Matz, E., Studier, F.W., Pegg, A.E., Coleman, C.S., Flanagan, J.M.(2006) Proc Natl Acad Sci U S A 103: 2063-2068
- PubMed: 16455797 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0511008103
- Primary Citation Related Structures:
2B3U, 2B3V, 2B4B, 2B4D, 2B58, 2B5G - PubMed Abstract:
Spermidine/spermine N1-acetyltransferase (SSAT) is a key enzyme in the control of polyamine levels in human cells, as acetylation of spermidine and spermine triggers export or degradation. Increased intracellular polyamine levels accompany several types of cancers as well as other human diseases, and compounds that affect the expression, activity, or stability of SSAT are being explored as potential therapeutic drugs. We have expressed human SSAT from the cloned cDNA in Escherichia coli and have determined high-resolution structures of wild-type and mutant SSAT, as the free dimer and in binary and ternary complexes with CoA, acetyl-CoA (AcCoA), spermine, and the inhibitor N1,N11bis-(ethyl)-norspermine (BE-3-3-3). These structures show details of binding sites for cofactor, substrates, and inhibitor and provide a framework to understand enzymatic activity, mutations, and the action of potential drugs. Two dimer conformations were observed: a symmetric form with two open surface channels capable of binding substrate or cofactor, and an asymmetric form in which only one of the surface channels appears capable of binding and acetylating polyamines. SSAT was found to self-acetylate lysine-26 in the presence of AcCoA and absence of substrate, a reaction apparently catalzyed by AcCoA bound in the second channel of the asymmetric dimer. These unexpected and intriguing complexities seem likely to have some as yet undefined role in regulating SSAT activity or stability as a part of polyamine homeostasis. Sequence signatures group SSAT with proteins that appear to have thialysine Nepsilon-acetyltransferase activity.
- Biology Department, Brookhaven National Laboratory, Upton, NY 11973, USA.
Organizational Affiliation: