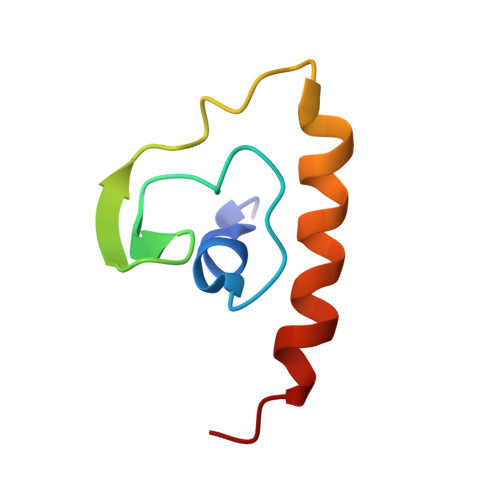
Solution structure of the N-terminal A domain of the human voltage-gated Ca2+channel beta4a subunit
Vendel, A.C., Rithner, C.D., Lyons, B.A., Horne, W.A.(2006) Protein Sci 15: 378-383
- PubMed: 16385006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051894506
- Primary Citation Related Structures:
2D46 - PubMed Abstract:
Ca2+ channel beta subunits regulate trafficking and gating (opening and closing) of voltage-dependent Ca2+ channel alpha1 subunits. Based on primary sequence comparisons, they are thought to be modular structures composed of five domains (A-E) that are related to the large family of membrane associated guanylate-kinase (MAGUK) proteins. The crystal structures of the beta subunit core, B-D, domains have recently been reported; however, very little is known about the structures of the A and E domains. The N-terminal A domain is a hypervariable region that differs among the four subtypes of Ca2+ channel beta subunits (beta1-beta4). Furthermore, this domain undergoes alternative splicing to create multiple N-terminal structures within a given gene class that have distinct effects on gating. We have solved the solution structure of the A domain of the human beta4a subunit, a splice variant that we have shown previously to have alpha1 subunit subtype-specific effects on Ca2+ channel trafficking and gating.
- Department of Biomedical Sciences, College of Veterinary Medicine and Biomedical Sciences, Colorado State University, Fort Collins, CO 80526, USA.
Organizational Affiliation: