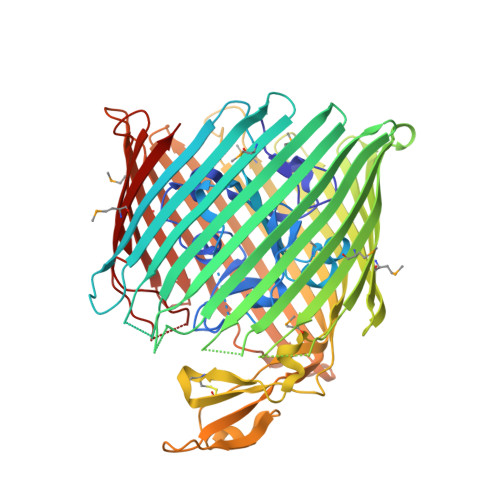
Structure of colicin I receptor bound to the R-domain of colicin Ia: implications for protein import.
Buchanan, S.K., Lukacik, P., Grizot, S., Ghirlando, R., Ali, M.M., Barnard, T.J., Jakes, K.S., Kienker, P.K., Esser, L.(2007) EMBO J 26: 2594-2604
- PubMed: 17464289 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601693
- Primary Citation Related Structures:
2HDF, 2HDI - PubMed Abstract:
Colicin Ia is a 69 kDa protein that kills susceptible Escherichia coli cells by binding to a specific receptor in the outer membrane, colicin I receptor (70 kDa), and subsequently translocating its channel forming domain across the periplasmic space, where it inserts into the inner membrane and forms a voltage-dependent ion channel. We determined crystal structures of colicin I receptor alone and in complex with the receptor binding domain of colicin Ia. The receptor undergoes large and unusual conformational changes upon colicin binding, opening at the cell surface and positioning the receptor binding domain of colicin Ia directly above it. We modelled the interaction with full-length colicin Ia to show that the channel forming domain is initially positioned 150 A above the cell surface. Functional data using full-length colicin Ia show that colicin I receptor is necessary for cell surface binding, and suggest that the receptor participates in translocation of colicin Ia across the outer membrane.
- Laboratory of Molecular Biology, National Institute of Diabetes & Digestive & Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA. skbuchan@helix.nih.gov
Organizational Affiliation: