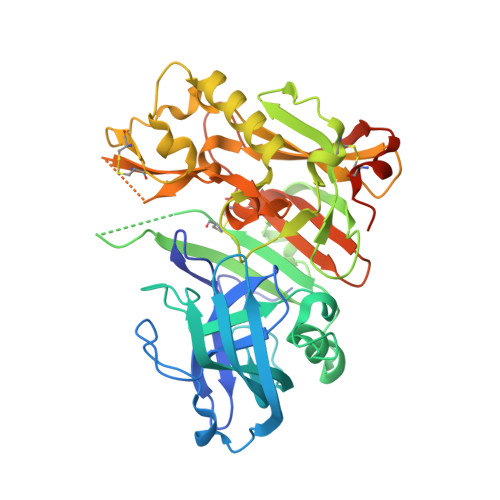
Design of potent inhibitors of human beta-secretase. Part 2.
Freskos, J.N., Fobian, Y.M., Benson, T.E., Moon, J.B., Bienkowski, M.J., Brown, D.L., Emmons, T.L., Heintz, R., Laborde, A., McDonald, J.J., Mischke, B.V., Molyneaux, J.M., Mullins, P.B., Bryan Prince, D., Paddock, D.J., Tomasselli, A.G., Winterrowd, G.(2007) Bioorg Med Chem Lett 17: 78-81
- PubMed: 17049233 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.09.091
- Primary Citation Related Structures:
2HM1 - PubMed Abstract:
We describe an optimized series of acyclic hydroxyethylamine transition state isosteres of beta-secretase that incorporates a variety of P(2) side chains that yield potent inhibitors with excellent cellular activity. A 2.2A crystal structure of compound 13 is shown.
- Pfizer Inc., 700N. Chesterfield Pkwy., St. Louis, MO 63198, USA.
Organizational Affiliation: