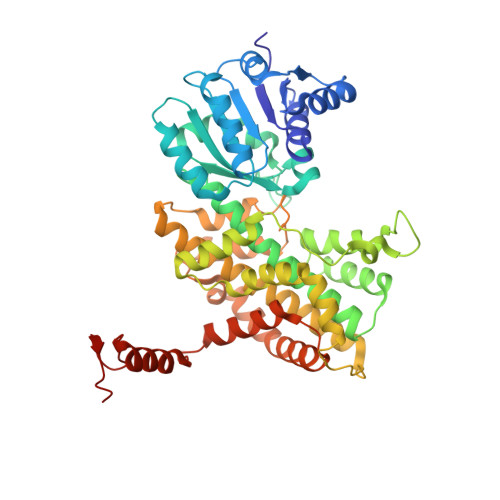
Crystal Structures of a Bacterial 6-Phosphogluconate Dehydrogenase Reveal Aspects of Specificity, Mechanism and Mode of Inhibition by Analogues of High-Energy Reaction Intermediates.
Sundaramoorthy, R., Iulek, J., Barrett, M.P., Bidet, O., Ruda, G.F., Gilbert, I.H., Hunter, W.N.(2007) FEBS J 274: 275
- PubMed: 17222187 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05585.x
- Primary Citation Related Structures:
2IYO, 2IYP, 2IZ0, 2IZ1 - PubMed Abstract:
Crystal structures of recombinant Lactococcus lactis 6-phosphogluconate dehydrogenase (LlPDH) in complex with substrate, cofactor, product and inhibitors have been determined. LlPDH shares significant sequence identity with the enzymes from sheep liver and the protozoan parasite Trypanosoma brucei for which structures have been reported. Comparisons indicate that the key residues in the active site are highly conserved, as are the interactions with the cofactor and the product ribulose 5-phosphate. However, there are differences in the conformation of the substrate 6-phosphogluconate which may reflect distinct states relevant to catalysis. Analysis of the complex formed with the potent inhibitor 4-phospho-d-erythronohydroxamic acid, suggests that this molecule does indeed mimic the high-energy intermediate state that it was designed to. The analysis also identified, as a contaminant by-product of the inhibitor synthesis, 4-phospho-d-erythronamide, which binds in similar fashion. LlPDH can now serve as a model system for structure-based inhibitor design targeting the enzyme from Trypanosoma species.
- Division of Biological Chemistry and Molecular Microbiology, College of Life Sciences, University of Dundee, UK.
Organizational Affiliation: