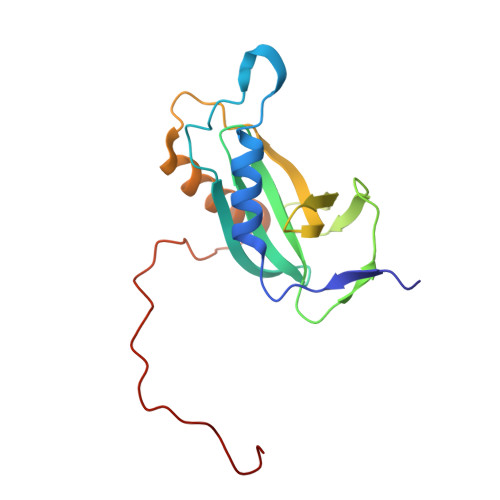
Solution structure of the Escherichia coli HybE reveals a novel fold
Shao, X., Lu, J., Hu, Y., Xia, B., Jin, C.(2009) Proteins 75: 1051-1056
- PubMed: 19291739 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22391
- Primary Citation Related Structures:
2KC5 - Beijing Nuclear Magnetic Resonance Center, Peking University, Beijing 100871, China.
Organizational Affiliation: