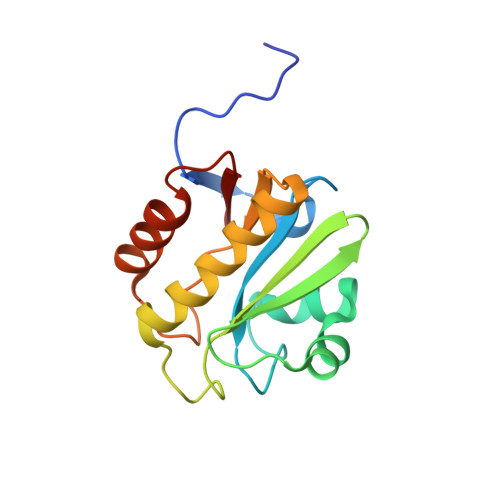
Orphan Macrodomain Protein (Human C6orf130) Is an O-Acyl-ADP-ribose Deacylase: SOLUTION STRUCTURE AND CATALYTIC PROPERTIES.
Peterson, F.C., Chen, D., Lytle, B.L., Rossi, M.N., Ahel, I., Denu, J.M., Volkman, B.F.(2011) J Biological Chem 286: 35955-35965
- PubMed: 21849506 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.276238
- Primary Citation Related Structures:
2L8R, 2LGR - PubMed Abstract:
Post-translational modification of proteins/histones by lysine acylation has profound effects on the physiological function of modified proteins. Deacylation by NAD(+)-dependent sirtuin reactions yields as a product O-acyl-ADP-ribose, which has been implicated as a signaling molecule in modulating cellular processes. Macrodomain-containing proteins are reported to bind NAD(+)-derived metabolites. Here, we describe the structure and function of an orphan macrodomain protein, human C6orf130. This unique 17-kDa protein is a stand-alone macrodomain protein that occupies a distinct branch in the phylogenic tree. We demonstrate that C6orf130 catalyzes the efficient deacylation of O-acetyl-ADP-ribose, O-propionyl-ADP-ribose, and O-butyryl-ADP-ribose to produce ADP-ribose (ADPr) and acetate, propionate, and butyrate, respectively. Using NMR spectroscopy, we solved the structure of C6orf130 in the presence and absence of ADPr. The structures showed a canonical fold with a deep ligand (ADPr)-binding cleft. Structural comparisons of apo-C6orf130 and the ADPr-C6orf130 complex revealed fluctuations of the β(5)-α(4) loop that covers the bound ADPr, suggesting that the β(5)-α(4) loop functions as a gate to sequester substrate and offer flexibility to accommodate alternative substrates. The ADPr-C6orf130 complex identified amino acid residues involved in substrate binding and suggested residues that function in catalysis. Site-specific mutagenesis and steady-state kinetic analyses revealed two critical catalytic residues, Ser-35 and Asp-125. We propose a catalytic mechanism for deacylation of O-acyl-ADP-ribose by C6orf130 and discuss the biological implications in the context of reversible protein acylation at lysine residues.
- Department of Biochemistry and Center for Eukaryotic Structural Genomics, Medical College of Wisconsin, Milwaukee, Wisconsin 53226.
Organizational Affiliation: