Structural insights into TAZ2 domain-mediated CBP/p300 recruitment by transactivation domain 1 of the lymphopoietic transcription factor E2A.
Lochhead, M.R., Brown, A.D., Kirlin, A.C., Chitayat, S., Munro, K., Findlay, J.E., Baillie, G.S., LeBrun, D.P., Langelaan, D.N., Smith, S.P.(2020) J Biological Chem 295: 4303-4315
- PubMed: 32098872 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.011078
- Primary Citation Related Structures:
2MH0 - PubMed Abstract:
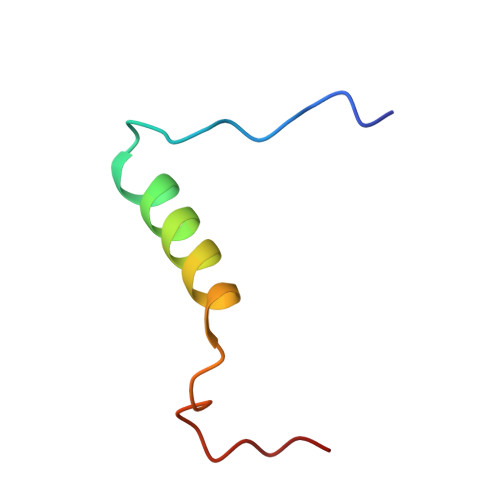
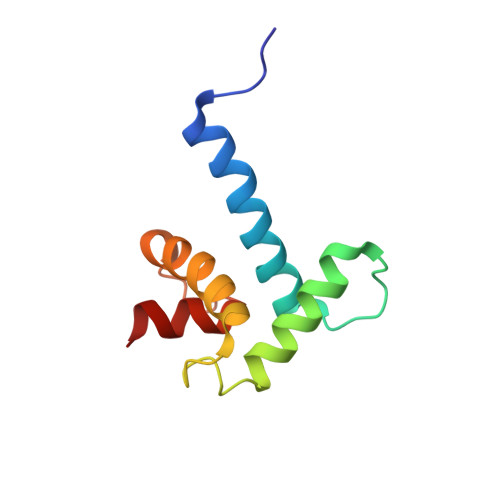
The E-protein transcription factors guide immune cell differentiation, with E12 and E47 (hereafter called E2A) being essential for B-cell specification and maturation. E2A and the oncogenic chimera E2A-PBX1 contain three transactivation domains (ADs), with AD1 and AD2 having redundant, independent, and cooperative functions in a cell-dependent manner. AD1 and AD2 both mediate their functions by binding to the KIX domain of the histone acetyltransferase paralogues CREB-binding protein (CBP) and E1A-binding protein P300 (p300). This interaction is necessary for B-cell maturation and oncogenesis by E2A-PBX1 and occurs through conserved Φ XX ΦΦ motifs (with Φ denoting a hydrophobic amino acid) in AD1 and AD2. However, disruption of this interaction via mutation of the KIX domain in CBP/p300 does not completely abrogate binding of E2A and E2A-PBX1. Here, we determined that E2A-AD1 and E2A-AD2 also interact with the TAZ2 domain of CBP/p300. Characterization of the TAZ2:E2A-AD1(1-37) complex indicated that E2A-AD1 adopts an α-helical structure and uses its Φ XX ΦΦ motif to bind TAZ2. Whereas this region overlapped with the KIX recognition region, key KIX-interacting E2A-AD1 residues were exposed, suggesting that E2A-AD1 could simultaneously bind both the KIX and TAZ2 domains. However, we did not detect a ternary complex involving E2A-AD1, KIX, and TAZ2 and found that E2A containing both intact AD1 and AD2 is required to bind to CBP/p300. Our findings highlight the structural plasticity and promiscuity of E2A-AD1 and suggest that E2A binds both the TAZ2 and KIX domains of CBP/p300 through AD1 and AD2.
- Department of Biomedical and Molecular Sciences, Queen's University, Kingston, Ontario K7L 3N6, Canada.
Organizational Affiliation: