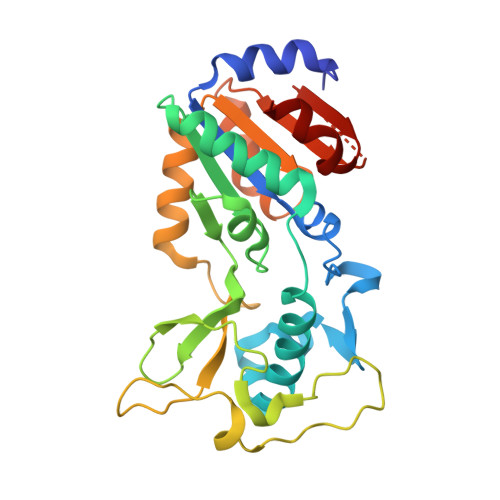
Structural basis of inhibition of the human NAD+-dependent deacetylase SIRT5 by suramin.
Schuetz, A., Min, J., Antoshenko, T., Wang, C.L., Allali-Hassani, A., Dong, A., Loppnau, P., Vedadi, M., Bochkarev, A., Sternglanz, R., Plotnikov, A.N.(2007) Structure 15: 377-389
- PubMed: 17355872 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2007.02.002
- Primary Citation Related Structures:
2NYR - PubMed Abstract:
Sirtuins are NAD(+)-dependent protein deacetylases and are emerging as molecular targets for the development of pharmaceuticals to treat human metabolic and neurological diseases and cancer. To date, several sirtuin inhibitors and activators have been identified, but the structural mechanisms of how these compounds modulate sirtuin activity have not yet been determined. We identified suramin as a compound that binds to human SIRT5 and showed that it inhibits SIRT5 NAD(+)-dependent deacetylase activity with an IC(50) value of 22 microM. To provide insights into how sirtuin function is altered by inhibitors, we determined two crystal structures of SIRT5, one in complex with ADP-ribose, the other bound to suramin. Our structural studies provide a view of a synthetic inhibitory compound in a sirtuin active site revealing that suramin binds into the NAD(+), the product, and the substrate-binding site. Finally, our structures may enable the rational design of more potent inhibitors.
- Structural Genomics Consortium, University of Toronto, 100 College Street, Toronto, Ontario M5G 1L5, Canada.
Organizational Affiliation: