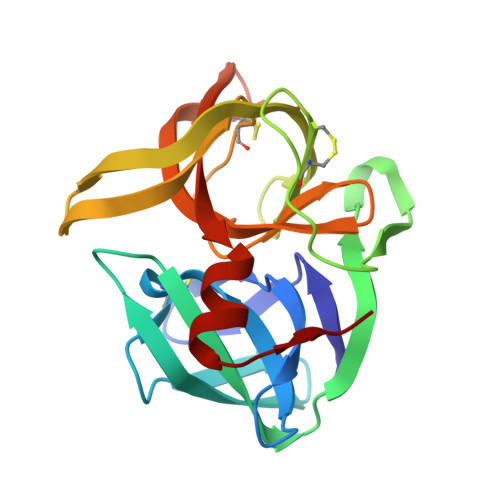
Structural and mechanistic exploration of Acid resistance: kinetic stability facilitates evolution of extremophilic behavior
Kelch, B.A., Eagen, K.P., Erciyas, F.P., Humphris, E.L., Thomason, A.R., Mitsuiki, S., Agard, D.A.(2007) J Mol Biology 368: 870-883
- PubMed: 17382344 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.02.032
- Primary Citation Related Structures:
2OUA - PubMed Abstract:
Kinetically stable proteins are unique in that their stability is determined solely by kinetic barriers rather than by thermodynamic equilibria. To better understand how kinetic stability promotes protein survival under extreme environmental conditions, we analyzed the unfolding behavior and determined the structure of Nocardiopsis alba Protease A (NAPase), an acid-resistant, kinetically stable protease, and compared these results with a neutrophilic homolog, alpha-lytic protease (alphaLP). Although NAPase and alphaLP have the same number of acid-titratable residues, kinetic studies revealed that the height of the unfolding free energy barrier for NAPase is less sensitive to acid than that of alphaLP, thereby accounting for NAPase's improved tolerance of low pH. A comparison of the alphaLP and NAPase structures identified multiple salt-bridges in the domain interface of alphaLP that were relocated to outer regions of NAPase, suggesting a novel mechanism of acid stability in which acid-sensitive electrostatic interactions are rearranged to similarly affect the energetics of both the native state and the unfolding transition state. An acid-stable variant of alphaLP in which a single interdomain salt-bridge is replaced with a corresponding intradomain NAPase salt-bridge shows a dramatic >15-fold increase in acid resistance, providing further evidence for this hypothesis. These observations also led to a general model of the unfolding transition state structure for alphaLP protease family members in which the two domains separate from each other while remaining relatively intact themselves. These results illustrate the remarkable utility of kinetic stability as an evolutionary tool for developing longevity over a broad range of harsh conditions.
- Howard Hughes Medical Institute and the Department of Biochemistry and Biophysics, University of California-San Francisco, 600 16th Street, San Francisco, CA 94158-2517, USA.
Organizational Affiliation: