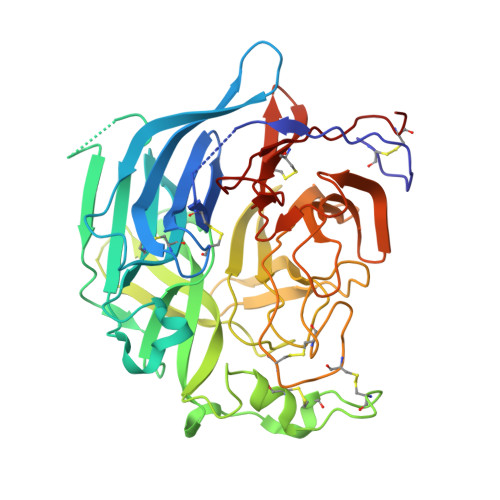
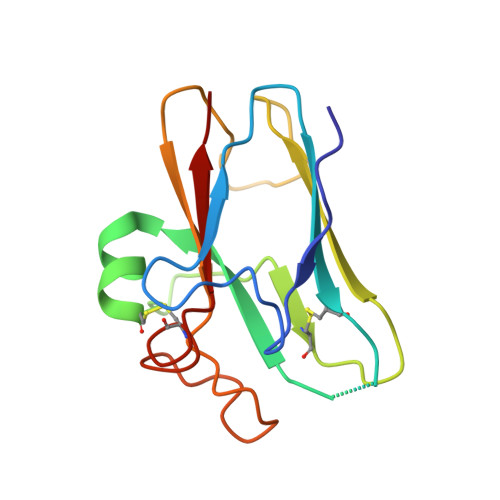
Structural Basis of Nipah and Hendra Virus Attachment to Their Cell-Surface Receptor Ephrin-B2
Bowden, T.A., Aricescu, A.R., Gilbert, R.J., Grimes, J.M., Jones, E.Y., Stuart, D.I.(2008) Nat Struct Mol Biol 15: 567
- PubMed: 18488039 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1435
- Primary Citation Related Structures:
2VSK, 2VSM - PubMed Abstract:
Nipah and Hendra viruses are emergent paramyxoviruses, causing disease characterized by rapid onset and high mortality rates, resulting in their classification as Biosafety Level 4 pathogens. Their attachment glycoproteins are essential for the recognition of the cell-surface receptors ephrin-B2 (EFNB2) and ephrin-B3 (EFNB3). Here we report crystal structures of both Nipah and Hendra attachment glycoproteins in complex with human EFNB2. In contrast to previously solved paramyxovirus attachment complexes, which are mediated by sialic acid interactions, the Nipah and Hendra complexes are maintained by an extensive protein-protein interface, including a crucial phenylalanine side chain on EFNB2 that fits snugly into a hydrophobic pocket on the viral protein. By analogy with the development of antivirals against sialic acid binding viruses, these results provide a structural template to target antiviral inhibition of protein-protein interactions.
- Division of Structural Biology, University of Oxford, Henry Wellcome Building of Genomic Medicine, Roosevelt Drive, Oxford OX3 7BN, UK.
Organizational Affiliation: