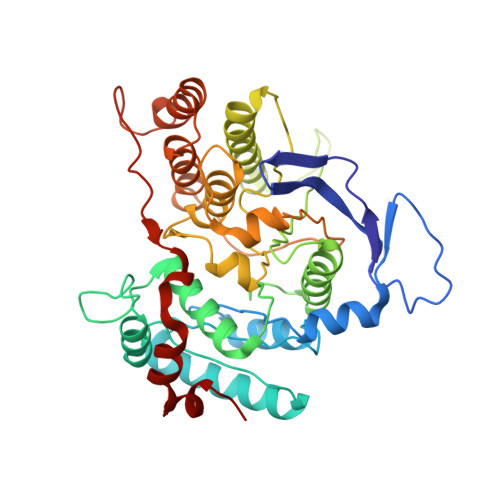
Structural Analysis of Thermus Thermophilus Hb27 Mannosyl-3-Phosphoglycerate Synthase Provides Evidence for a Second Catalytic Metal Ion and New Insight Into the Retaining Mechanism of Glycosyltransferases.
Goncalves, S., Borges, N., Esteves, A.M., Victor, B., Soares, C.M., Santos, H., Matias, P.M.(2010) J Biological Chem 285: 17857
- PubMed: 20356840 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.095976
- Primary Citation Related Structures:
2WVK, 2WVL, 2WVM - PubMed Abstract:
Mannosyl-3-phosphoglycerate synthase is a glycosyltransferase involved in the two-step synthetic pathway of mannosylglycerate, a compatible solute that accumulates in response to salt and/or heat stresses in many microorganisms thriving in hot environments. The three-dimensional structure of mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 in its binary complex form, with GDP-alpha-D-mannose and Mg(2+), shows a second metal binding site, about 6 A away from the mannose moiety. Kinetic and mutagenesis studies have shown that this metal site plays a role in catalysis. Additionally, Asp(167) in the DXD motif is found within van der Waals contact distance of the C1' atom in the mannopyranose ring, suggesting its action as a catalytic nucleophile, either in the formation of a glycosyl-enzyme intermediate according to the double-displacement S(N)2 reaction mechanism or in the stabilization of the oxocarbenium ion-like intermediate according to the D(N)*A(Nss) (S(N)i-like) reaction mechanism. We propose that either mechanism may occur in retaining glycosyltransferases with a GT-A fold, and, based on the gathered structural information, we identified an extended structural signature toward a common scaffold between the inverting and retaining glycosyltransferases.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Apartado 127, 2781-901 Oeiras, Portugal.
Organizational Affiliation: