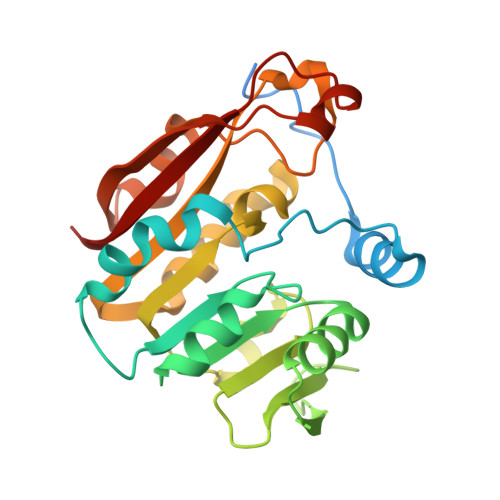
Structural Basis of Substrate Recognition in Thiopurine S-Methyltransferase
Peng, Y., Feng, Q., Wilk, D., Adjei, A.A., Salavaggione, O.E., Weinshilboum, R.M., Yee, V.C.(2008) Biochemistry 47 (23), 6216 6225, 2008. 10.1021/bi800102x: 6216-6225
- PubMed: 18484748 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi800102x
- Primary Citation Related Structures:
3BGD, 3BGI - PubMed Abstract:
Thiopurine S-methyltransferase (TPMT) modulates the cytotoxic effects of thiopurine prodrugs such as 6-mercaptopurine by methylating them in a reaction using S-adenosyl- l-methionine as the donor. Patients with TPMT variant allozymes exhibit diminished levels of protein and/or enzyme activity and are at risk for thiopurine drug-induced toxicity. We have determined two crystal structures of murine TPMT, as a binary complex with the product S-adenosyl- l-homocysteine and as a ternary complex with S-adenosyl- l-homocysteine and the substrate 6-mercaptopurine, to 1.8 and 2.0 A resolution, respectively. Comparison of the structures reveals that an active site loop becomes ordered upon 6-mercaptopurine binding. The positions of the two ligands are consistent with the expected S N2 reaction mechanism. Arg147 and Arg221, the only polar amino acids near 6-mercaptopurine, are highlighted as possible participants in substrate deprotonation. To probe whether these residues are important for catalysis, point mutants were prepared in the human enzyme. Substitution of Arg152 (Arg147 in murine TPMT) with glutamic acid decreases V max and increases K m for 6-mercaptopurine but not K m for S-adenosyl- l-methionine. Substitution at this position with alanine or histidine and similar substitutions of Arg226 (Arg221 in murine TPMT) result in no effect on enzyme activity. The double mutant Arg152Ala/Arg226Ala exhibits a decreased V max and increased K m for 6-mercaptopurine. These observations suggest that either Arg152 or Arg226 may participate in some fashion in the TPMT reaction, with one residue compensating when the other is altered, and that Arg152 may interact with substrate more directly than Arg226, consistent with observations in the murine TPMT crystal structure.
- Department of Biochemistry, Case Western Reserve University, Cleveland, Ohio 44106, USA.
Organizational Affiliation: