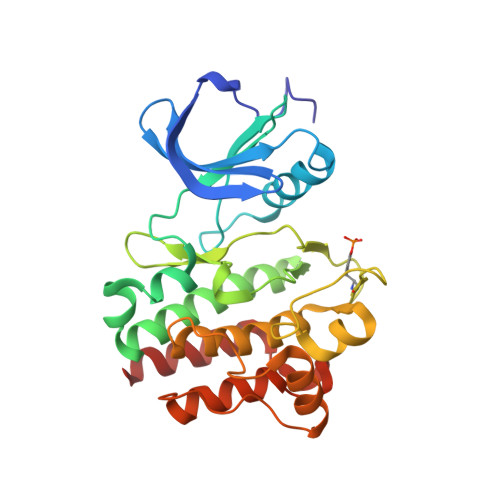
Structure-based design of novel 2-amino-6-phenyl-pyrimido[5',4':5,6]pyrimido[1,2-a]benzimidazol-5(6H)-ones as potent and orally active inhibitors of lymphocyte specific kinase (Lck): synthesis, SAR, and in vivo anti-inflammatory activity.
Martin, M.W., Newcomb, J., Nunes, J.J., Boucher, C., Chai, L., Epstein, L.F., Faust, T., Flores, S., Gallant, P., Gore, A., Gu, Y., Hsieh, F., Huang, X., Kim, J.L., Middleton, S., Morgenstern, K., Oliveira-dos-Santos, A., Patel, V.F., Powers, D., Rose, P., Tudor, Y., Turci, S.M., Welcher, A.A., Zack, D., Zhao, H., Zhu, L., Zhu, X., Ghiron, C., Ermann, M., Johnston, D., Saluste, C.G.(2008) J Med Chem 51: 1637-1648
- PubMed: 18278858 Search on PubMed
- DOI: https://doi.org/10.1021/jm701095m
- Primary Citation Related Structures:
3BYM, 3BYO - PubMed Abstract:
Lck, or lymphocyte specific kinase, is a cytoplasmic tyrosine kinase of the Src family expressed in T-cells and NK cells. Genetic evidence from knockout mice and human mutations demonstrates that Lck kinase activity is critical for T-cell receptor-mediated signaling, leading to normal T-cell development and activation. A small molecule inhibitor of Lck is expected to be useful in the treatment of T-cell-mediated autoimmune and inflammatory disorders and/or organ transplant rejection. In this paper, we describe the structure-guided design, synthesis, structure-activity relationships, and pharmacological characterization of 2-amino-6-phenylpyrimido[5',4':5,6]pyrimido[1,2- a]benzimidazol-5(6 H)-ones, a new class of compounds that are potent inhibitors of Lck. The most promising compound of this series, 6-(2,6-dimethylphenyl)-2-((4-(4-methyl-1-piperazinyl)phenyl)amino)pyrimido[5',4':5,6]pyrimido-[1,2- a]benzimidazol-5(6 H)-one ( 25), exhibits potent inhibition of Lck kinase activity. This activity translates into inhibition of in vitro cell-based assays and in vivo models of T-cell activation and arthritis, respectively.
- Department of Medicinal Chemistry, Amgen Inc., One Kendall Square, Cambridge, MA 02139, USA. matmarti@amgen.com
Organizational Affiliation: