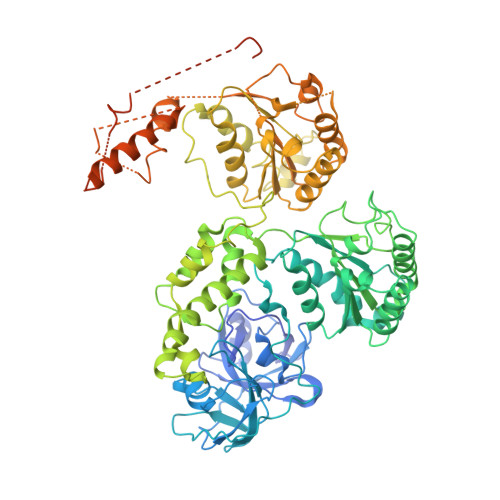
Improved structures of full-length p97, an AAA ATPase: implications for mechanisms of nucleotide-dependent conformational change.
Davies, J.M., Brunger, A.T., Weis, W.I.(2008) Structure 16: 715-726
- PubMed: 18462676 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2008.02.010
- Primary Citation Related Structures:
3CF0, 3CF1, 3CF2, 3CF3 - PubMed Abstract:
The ATPases associated with various cellular activities (AAA) protein p97 has been implicated in a variety of cellular processes, including endoplasmic reticulum-associated degradation and homotypic membrane fusion. p97 belongs to a subgroup of AAA proteins that contains two nucleotide binding domains, D1 and D2. We determined the crystal structure of D2 at 3.0 A resolution. This model enabled rerefinement of full-length p97 in different nucleotide states against previously reported low-resolution diffraction data to significantly improved R values and Ramachandran statistics. Although the overall fold remained similar, there are significant improvements, especially around the D2 nucleotide binding site. The rerefinement illustrates the importance of knowledge of high-resolution structures of fragments covering most of the whole molecule. The structures suggest that nucleotide hydrolysis is transformed into larger conformational changes by pushing of one D2 domain by its neighbor in the hexamer, and transmission of nucleotide-state information through the D1-D2 linker to displace the N-terminal, effector binding domain.
- Department of Structural Biology, Stanford University, Stanford, CA 94305-5432, USA.
Organizational Affiliation: