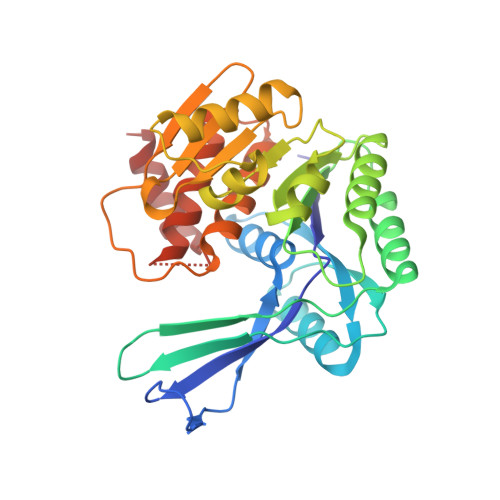
Crystallographic structure of phosphofructokinase-2 from Escherichia coli in complex with two ATP molecules. Implications for substrate inhibition.
Cabrera, R., Ambrosio, A.L., Garratt, R.C., Guixe, V., Babul, J.(2008) J Mol Biology 383: 588-602
- PubMed: 18762190 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.08.029
- Primary Citation Related Structures:
3CQD - PubMed Abstract:
Phosphofructokinase-1 and -2 (Pfk-1 and Pfk-2, respectively) from Escherichia coli belong to different homologous superfamilies. However, in spite of the lack of a common ancestor, they share the ability to catalyze the same reaction and are inhibited by the substrate MgATP. Pfk-2, an ATP-dependent 6-phosphofructokinase member of the ribokinase-like superfamily, is a homodimer of 66 kDa subunits whose oligomerization state is necessary for catalysis and stability. The presence of MgATP favors the tetrameric form of the enzyme. In this work, we describe the structure of Pfk-2 in its inhibited tetrameric form, with each subunit bound to two ATP molecules and two Mg ions. The present structure indicates that substrate inhibition occurs due to the sequential binding of two MgATP molecules per subunit, the first at the usual site occupied by the nucleotide in homologous enzymes and the second at the allosteric site, making a number of direct and Mg-mediated interactions with the first. Two configurations are observed for the second MgATP, one of which involves interactions with Tyr23 from the adjacent subunit in the dimer and the other making an unusual non-Watson-Crick base pairing with the adenine in the substrate ATP. The oligomeric state observed in the crystal is tetrameric, and some of the structural elements involved in the binding of the substrate and allosteric ATPs are also participating in the dimer-dimer interface. This structure also provides the grounds to compare analogous features of the nonhomologous phosphofructokinases from E. coli.
- Departamento de Biología, Facultad de Ciencias, Universidad de Chile, Casilla 653, Santiago, Chile.
Organizational Affiliation: