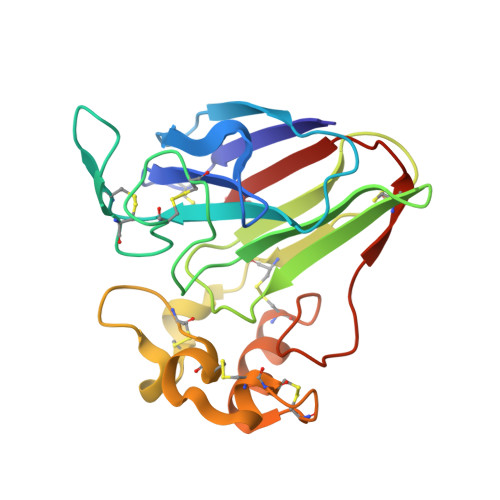
A magic triangle for experimental phasing of macromolecules
Beck, T., Krasauskas, A., Gruene, T., Sheldrick, G.M.(2008) Acta Crystallogr D Biol Crystallogr 64: 1179-1182
- PubMed: 19020357 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908030266
- Primary Citation Related Structures:
3E3D, 3E3S, 3E3T - PubMed Abstract:
Obtaining phase information for the solution of macromolecular structures is still one of the bottlenecks in X-ray crystallography. 5-Amino-2,4,6-triiodoisophthalic acid (I3C), in which three covalently bound iodines form an equilateral triangle, was incorporated into proteins in order to obtain phases by single-wavelength anomalous dispersion (SAD). An improved binding capability compared with simple heavy-metal ions, ready availability, improved recognition of potential heavy-atom sites and low toxicity make I3C particularly suitable for experimental phasing.
- Department of Structural Chemistry, Georg-August-Universität Göttingen, Tammannstrasse 4, 37077 Göttingen, Germany. tbeck@shelx.uni-ac.gwdg.de
Organizational Affiliation: